 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT5G46350.1 | ||||||||
| Common Name | ATWRKY8, MPL12.15, WRKY8 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 326aa MW: 37294 Da PI: 8.0888 | ||||||||
| Description | WRKY DNA-binding protein 8 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 104.2 | 7.1e-33 | 183 | 240 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
+Dgy+WrKYGqK vk+s++prsYYrCt+++C+vkk+vers +dp+vv++tYe++Hnh+
AT5G46350.1 183 EDGYRWRKYGQKAVKNSPYPRSYYRCTTQKCNVKKRVERSYQDPTVVITTYESQHNHP 240
8********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF038130 | 1.2E-112 | 1 | 326 | IPR017396 | PWRKY transcription factor, group IIc |
| Gene3D | G3DSA:2.20.25.80 | 3.2E-34 | 167 | 242 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 5.62E-29 | 174 | 242 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 32.282 | 177 | 242 | IPR003657 | WRKY domain |
| SMART | SM00774 | 6.1E-38 | 182 | 241 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 2.1E-25 | 183 | 240 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0051607 | Biological Process | defense response to virus | ||||
| GO:0070301 | Biological Process | cellular response to hydrogen peroxide | ||||
| GO:1901002 | Biological Process | positive regulation of response to salt stress | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025195 | anatomy | pollen tube cell | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 326 aa Download sequence Send to blast |
MSHEIKDLNN YHYTSSYNHY NINNQNMINL PYVSGPSAYN ANMISSSQVG FDLPSKNLSP 60 QGAFELGFEL SPSSSDFFNP SLDQENGLYN AYNYNSSQKS HEVVGDGCAT IKSEVRVSAS 120 PSSSEADHHP GEDSGKIRKK REVRDGGEDD QRSQKVVKTK KKEEKKKEPR VSFMTKTEVD 180 HLEDGYRWRK YGQKAVKNSP YPRSYYRCTT QKCNVKKRVE RSYQDPTVVI TTYESQHNHP 240 IPTNRRTAMF SGTTASDYNP SSSPIFSDLI INTPRSFSND DLFRVPYASV NVNPSYHQQQ 300 HGFHQQESEF ELLKEMFPSV FFKQEP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wj2_A | 4e-28 | 172 | 242 | 7 | 77 | Probable WRKY transcription factor 4 |
| 2lex_A | 4e-28 | 172 | 242 | 7 | 77 | Probable WRKY transcription factor 4 |
| Search in ModeBase | ||||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| Genevisible | 248896_at | 0.0 | ||||
| Expression Atlas | AT5G46350 | - | ||||
| AtGenExpress | AT5G46350 | - | ||||
| ATTED-II | AT5G46350 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in roots and at lower levels in rosette leaves, cauline leaves, stems, flowers and siliques. {ECO:0000269|PubMed:23451802}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | member of WRKY Transcription Factor; Group II-c | |||||
| UniProt | Transcription factor. Interacts specifically with the W box (5'-TTGAC[CT]-3'), a frequently occurring stress-responsive cis-acting element. Functions as positive regulator of salt stress response. Binds the W box of LTI78/RD29A stress-response gene and directly regulates its transcription under salt stress. Functions antagonistically with VQ9 to regulate sodium and potassium homeostasis under salt stress by regulating the expression of downstream SOS (SALT OVERLY SENSITIVE) stress-responsive genes. The DNA-binding activity of WRKY8 is decreased by VQ9 (PubMed:23451802). Functions as negative regulator of basal resistance to the bacterial pathogen P. syringae and as positive regulator of resistance to the fungal pathogen to B. cinerea (PubMed:20367464). Functions as positive regulator of defense response againt tobamovirus (TMV) by regulating both the abscisic acid and ethylene signaling pathways. Positively regulates ABI4 expression and negatively modulates ACS6 and ERF104 expression by directly binding to the W box consensus motifs within their promoters (PubMed:23650359). {ECO:0000269|PubMed:20367464, ECO:0000269|PubMed:23451802, ECO:0000269|PubMed:23650359}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00545 | DAP | 27203113 | Download |
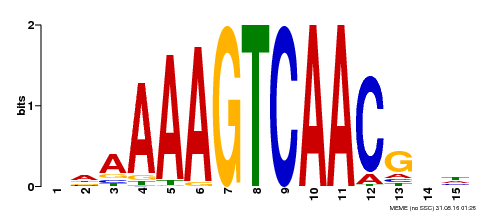
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT5G46350.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By wounding, abscisic acid (ABA), salicylic acid (SA), H(2)O(2), and infection with P.syringae pv. tomato DC3000 and B.cinerea (PubMed:20367464). Induced by salt stress (PubMed:23451802). {ECO:0000269|PubMed:20367464, ECO:0000269|PubMed:23451802}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Regulation -- Hormone ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hormone | |||||
| AHD | salicylic acid | |||||
| Phenotype -- Disruption Phenotype ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | DISRUPTION PHENOTYPE: No visible phenotype under normal growth conditions (PubMed:20367464, PubMed:23451802). Mutant plants show increased resistance to the bacterial pathogen P. syringae and enhanced susceptibility to the fungal pathogen to B. cinerea (PubMed:20367464). Mutant plants display increased sensitiviy to salt stress (PubMed:23451802). {ECO:0000269|PubMed:20367464, ECO:0000269|PubMed:23451802}. | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT5G46350 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF404855 | 0.0 | AF404855.1 Arabidopsis thaliana WRKY transcription factor 8 (WRKY8) mRNA, complete cds. | |||
| GenBank | AY063916 | 0.0 | AY063916.1 Arabidopsis thaliana unknown protein (At5g46350) mRNA, complete cds. | |||
| GenBank | AY096426 | 0.0 | AY096426.1 Arabidopsis thaliana unknown protein (At5g46350) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_199447.1 | 0.0 | WRKY DNA-binding protein 8 | ||||
| Swissprot | Q9FL26 | 0.0 | WRKY8_ARATH; WRKY transcription factor 8 | ||||
| TrEMBL | A0A178U9B9 | 0.0 | A0A178U9B9_ARATH; WRKY transcription factor | ||||
| STRING | AT5G46350.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1524 | 26 | 93 | Representative plant | OGRP14 | 17 | 875 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT5G46350.1 |
| Entrez Gene | 834678 |
| iHOP | AT5G46350 |
| wikigenes | AT5G46350 |



