 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Thhalv10000027m | ||||||||
| Common Name | EUTSA_v10000026mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Eutremeae; Eutrema
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1041aa MW: 116729 Da PI: 5.1994 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 181.7 | 8.3e-57 | 20 | 136 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfq 95
+e ++rwl++ ei++iL+n++k+++++e++t+p+sgs++L++rk++ryfrkDG++w+kk+dgktv+E+he+LK g+v+vl+cyYah+++n++fq
Thhalv10000027m 20 SEaRNRWLRPPEICEILQNYQKFQISTEPPTTPSSGSVFLFDRKVLRYFRKDGHNWRKKRDGKTVKEAHERLKAGSVDVLHCYYAHGQDNENFQ 113
4449****************************************************************************************** PP
CG-1 96 rrcywlLeeelekivlvhylevk 118
rr+yw+L+eel++iv+vhylevk
Thhalv10000027m 114 RRSYWMLQEELSHIVFVHYLEVK 136
********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.949 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.7E-84 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.2E-49 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 7.5E-5 | 464 | 552 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 6.65E-17 | 466 | 551 | IPR014756 | Immunoglobulin E-set |
| CDD | cd00204 | 6.67E-11 | 628 | 758 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 6.1E-15 | 629 | 761 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 16.812 | 632 | 758 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 4.04E-15 | 644 | 761 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.683 | 699 | 731 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 2.12E-8 | 859 | 911 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.92 | 860 | 882 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.029 | 862 | 890 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0042 | 863 | 881 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0011 | 883 | 905 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.468 | 884 | 908 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 1.2E-4 | 886 | 905 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1041 aa Download sequence Send to blast |
MAEARRFSPN NELDVGQILS EARNRWLRPP EICEILQNYQ KFQISTEPPT TPSSGSVFLF 60 DRKVLRYFRK DGHNWRKKRD GKTVKEAHER LKAGSVDVLH CYYAHGQDNE NFQRRSYWML 120 QEELSHIVFV HYLEVKGSRV STSYNRMQRT EDAARSPQET GEALTSERDG YASCSINQYD 180 HSNHSQATDS ASVNNVHTPE LEDAESAYNQ QPSSIIYSHQ ELQQPATERA NTGFDSYYQM 240 SLTPRDSYQK DLRVIPVTNS SNMIDKSRTI NGPGVTNGLR SKKSIDSQTW EEILGNCGSG 300 AEGLPMQPNS EHEVLDQILQ SCSFTMQDFA SLQESMVKSQ NQELNSGPTS DRTMWFQGQD 360 IELNAISNLA SNEKAPYLST MKQHLLDGAL GEEGLKKMDS FNRWMSKELG DVGVIANANE 420 SFTHSSSTAY WEEVDSENGS NGHNSRRDLD GYVMSPSLAK EQLFSITDFA PSWTYVGCEV 480 QVLVTGKFLK TREEAEMREW SCMFGQTEVP AEVIANGVLQ CVAPMHEAGR VPFYVSCSNR 540 LACSEVREFE YKVVEAPVLD GETDESTTCN TIEGLEARFI KLLCSKSDNP SSSLSGNDSD 600 LSQVSEKISL LLFENDDQLD QMLMNEISQE NMKNNLLQEA LKESLHSWLL QKIAEGGKGP 660 NVLDEGGQGI LHFAAALGYN WALEPTIVAG VSVDFRDVNG WTALHWAAFF GRELIIGSLI 720 ALGAAPGTLT DPNPDFPSGS TPSDLAYANG YKGIAGYLSE YALRAHVSLL SLNDNNAETS 780 LAAVETAPSQ SSSSLTDSLT AVRNATQAAA RIHQVFRAQS FQKKQIKEFG DRKFGMSEER 840 ALSMLAPKTH KPGRAHSDDS VQAAAIRIQN KFRGYKGRKD YLITRQRIIK IQAHVRGYQV 900 RKNYRKIIWS VGILEKVILR WRRKGAGLRG FKSDALVDKM QDGTEKEEDD DFFKQGRKQT 960 EERLQKALAR VKSMVQYPEA RDQYRRLLNV VNDIQESKVE KALENSEEAT CFDDDLIDIE 1020 ALLQDDDTLM LPMSSTLWNP * |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
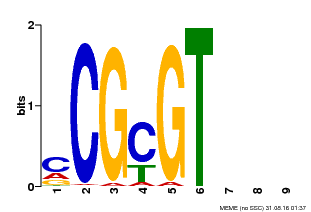
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Thhalv10000027m |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF506697 | 0.0 | AF506697.1 Arabidopsis thaliana calmodulin-binding transcription factor SR1 (SR1) mRNA, complete cds. | |||
| GenBank | AY510025 | 0.0 | AY510025.1 Arabidopsis thaliana ethylene-induced calmodulin-binding protein 1 mRNA, complete cds. | |||
| GenBank | BT002459 | 0.0 | BT002459.1 Arabidopsis thaliana Unknown protein mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024014331.1 | 0.0 | calmodulin-binding transcription activator 3 isoform X2 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | V4LQS4 | 0.0 | V4LQS4_EUTSA; Uncharacterized protein | ||||
| STRING | XP_006404687.1 | 0.0 | (Eutrema salsugineum) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Thhalv10000027m |
| Entrez Gene | 18021748 |



