 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010929993.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | C3H | ||||||||
| Protein Properties | Length: 499aa MW: 54497 Da PI: 7.3285 | ||||||||
| Description | C3H family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-CCCH | 32.9 | 1.1e-10 | 104 | 126 | 4 | 26 |
---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 4 elCrffartGtCkyGdrCkFaHg 26
+ C ++ ++GtC++G++CkF+H+
XP_010929993.1 104 PDCTYYVKNGTCRFGSSCKFNHP 126
68********************8 PP
| |||||||
| 2 | zf-CCCH | 22.6 | 1.8e-07 | 220 | 242 | 5 | 27 |
--SGGGGTS--TTTTT-SS-SSS CS
zf-CCCH 5 lCrffartGtCkyGdrCkFaHgp 27
+C++++ G Ck+G Ck+ H p
XP_010929993.1 220 ECKYYSMPGGCKFGKACKYVHHP 242
6********************75 PP
| |||||||
| 3 | zf-CCCH | 31.8 | 2.4e-10 | 261 | 286 | 1 | 26 |
--S---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 1 yktelCrffartGtCkyGdrCkFaHg 26
+++++C++++rtG Ck+ +Ck++H+
XP_010929993.1 261 PGERECPYYMRTGSCKFATNCKYHHP 286
5789*********************9 PP
| |||||||
| 4 | zf-CCCH | 24.9 | 3.5e-08 | 403 | 429 | 1 | 27 |
--S---SGGGGTS--TTTTT-SS-SSS CS
zf-CCCH 1 yktelCrffartGtCkyGdrCkFaHgp 27
+++++C++f++ G Ck+ CkF+H++
XP_010929993.1 403 PGQPECQYFMKHGVCKFKTACKFHHPK 429
5789*********************96 PP
| |||||||
| 5 | zf-CCCH | 35.4 | 1.8e-11 | 450 | 474 | 2 | 26 |
-S---SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 2 ktelCrffartGtCkyGdrCkFaHg 26
+++ C ++ r+G CkyG++CkF H+
XP_010929993.1 450 DQPVCTHYNRFGVCKYGPSCKFDHP 474
6899********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50103 | 14.643 | 100 | 128 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 9.0E-5 | 100 | 127 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 7.2E-6 | 101 | 128 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 2.8E-8 | 104 | 126 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 4.1E-4 | 106 | 126 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 12.54 | 215 | 243 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 0.067 | 215 | 242 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 7.72E-5 | 219 | 245 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 5.0E-5 | 220 | 241 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 3.7E-4 | 221 | 242 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 3.2E-5 | 260 | 287 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 14.596 | 260 | 288 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 3.3E-7 | 262 | 286 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 1.83E-5 | 263 | 287 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 13.704 | 402 | 430 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 0.0013 | 402 | 429 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 1.8E-5 | 404 | 429 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 5.36E-5 | 406 | 430 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 1.3E-4 | 448 | 475 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 15.403 | 448 | 476 | IPR000571 | Zinc finger, CCCH-type |
| SuperFamily | SSF90229 | 1.09E-5 | 449 | 475 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 1.1E-8 | 450 | 474 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 1.0E-4 | 453 | 473 | IPR000571 | Zinc finger, CCCH-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 499 aa Download sequence Send to blast |
MAVSEAEVVP KTEELGLGLA TAASSPSSPS DSFVEALTGE FASLKLEKKS AEEAGDQDNT 60 GAGDQEEITG DENGGVVEKD SGSPGKEKEG KEERFRYPQR LDAPDCTYYV KNGTCRFGSS 120 CKFNHPARRK RNKVKSKKPF EANQAVKEEQ KEPIPKVGQT ESKVDKKEEK ASSDMVGQME 180 CEAVDEMQNG TLSERGEEKD SKATEEKGKE TLSERAGQME CKYYSMPGGC KFGKACKYVH 240 HPGKLENNSV QLNFLGLPIR PGERECPYYM RTGSCKFATN CKYHHPDPTA VGGQDPRSGH 300 RNSEPPQHGL GAPQVPMTWP VQRMSNEPIS FLNASPSYVP GMILPPQGFH SNPEWNGYQA 360 PVSPLFPPDI HARNSASTIN HKTNKGDIPV SQQVPVDEYP ERPGQPECQY FMKHGVCKFK 420 TACKFHHPKT RPQTVPVGVL SPVGLPLKPD QPVCTHYNRF GVCKYGPSCK FDHPIGHGSS 480 ASAGVVVSGQ IATPNNATT |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 127 | 137 | RRKRNKVKSKK |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
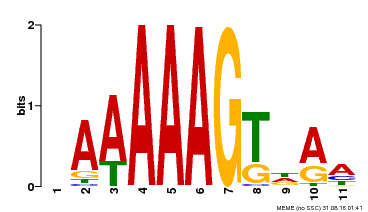
| Motif ID | Method | Source | Motif file |
| MP00582 | DAP | Transfer from AT5G63260 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010929993.1 | 0.0 | zinc finger CCCH domain-containing protein 65 isoform X1 | ||||
| TrEMBL | A0A2H3YAU2 | 0.0 | A0A2H3YAU2_PHODC; zinc finger CCCH domain-containing protein 65 isoform X1 | ||||
| STRING | XP_008796043.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7928 | 35 | 48 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G63260.2 | 1e-73 | C3H family protein | ||||



