 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | XP_010913727.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Arecoideae; Cocoseae; Elaeidinae; Elaeis
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 330aa MW: 35508.5 Da PI: 6.9634 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 50 | 4.1e-16 | 250 | 285 | 1 | 34 |
GATA 1 CsnCgttk..TplWRrgpdgnktLCnaCGlyyrkkg 34
C++Cg + Tp++Rrgpdg++tLCnaCGl ++ kg
XP_010913727.1 250 CHHCGISAksTPMMRRGPDGPRTLCNACGLVWANKG 285
****977777***********************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51320 | 13.959 | 114 | 149 | IPR010399 | Tify domain |
| SMART | SM00979 | 3.3E-11 | 114 | 149 | IPR010399 | Tify domain |
| Pfam | PF06200 | 1.4E-12 | 118 | 148 | IPR010399 | Tify domain |
| PROSITE profile | PS51017 | 13.344 | 175 | 217 | IPR010402 | CCT domain |
| Pfam | PF06203 | 3.1E-16 | 175 | 217 | IPR010402 | CCT domain |
| SMART | SM00401 | 2.6E-13 | 244 | 297 | IPR000679 | Zinc finger, GATA-type |
| PROSITE profile | PS50114 | 9.447 | 244 | 306 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 1.05E-11 | 248 | 295 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 9.0E-16 | 249 | 291 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 3.34E-16 | 249 | 300 | No hit | No description |
| Pfam | PF00320 | 7.7E-14 | 250 | 285 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 250 | 277 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 330 aa Download sequence Send to blast |
MSENPNPPVD VKPAAGGRFA GHHIHKPVAA HHVLPVVAMP AQIDDGGGGG GESQLIAAAD 60 AQALRQYEEA EHHSGVGVHG MEEDRDVGGD GEGMEADGQV DPSNMGDPHA VIVHQVGGNQ 120 LTLSFQGEVY VFDSVSPEKV QAVLLLLGER EMNAALNSFP SSSHMHNKRT NFPHRVASLM 180 RFREKRKERN FDKRIRYNVR KEVALRMQRN KGQFISSKSK PEDSTSGVTS WDTTQQWGSQ 240 ENRPPAASAC HHCGISAKST PMMRRGPDGP RTLCNACGLV WANKGIMRDL SKNPSPAIPN 300 ALSEPREGNN SAESGAAEPL AMTANGHDSS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. {ECO:0000250|UniProtKB:Q8LAU9}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00369 | DAP | Transfer from AT3G21175 | Download |
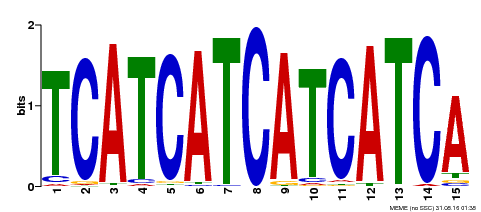
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By abscisic acid (ABA), and drought and salt stresses. Down-regulated by jasmonate and wounding. {ECO:0000269|PubMed:19618278}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010913727.1 | 0.0 | GATA transcription factor 17 | ||||
| Swissprot | Q5Z4U5 | 1e-101 | GAT20_ORYSJ; GATA transcription factor 20 | ||||
| TrEMBL | A0A2H3XCJ1 | 0.0 | A0A2H3XCJ1_PHODC; GATA transcription factor 20-like isoform X1 | ||||
| STRING | XP_008781787.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4325 | 38 | 68 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G21175.2 | 1e-80 | ZIM-like 1 | ||||



