 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Eucgr.K02763.1.p | ||||||||
| Common Name | EUGRSUZ_K02763 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1213aa MW: 135846 Da PI: 9.0799 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 11 | 0.0013 | 1097 | 1122 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
ykC C++sF +k +L+ H ++ +
Eucgr.K02763.1.p 1097 YKCDleGCTMSFGSKQELTLHKKNvC 1122
99*******************99866 PP
| |||||||
| 2 | zf-C2H2 | 13.7 | 0.00019 | 1121 | 1144 | 2 | 23 |
EET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCp..dCgksFsrksnLkrHirtH 23
+Cp Cgk F ++ +L++H r+H
Eucgr.K02763.1.p 1121 VCPvkGCGKKFFSHKYLVQHRRVH 1144
69999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 11.2 | 0.0012 | 1180 | 1206 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y C Cg++F+ s++ rH r+ H
Eucgr.K02763.1.p 1180 YICAepGCGQTFRFVSDFSRHKRKtgH 1206
789999****************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00558 | 4.3E-46 | 3 | 172 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 33.213 | 6 | 172 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 5.77E-25 | 18 | 187 | No hit | No description |
| Pfam | PF02373 | 1.1E-36 | 36 | 155 | IPR003347 | JmjC domain |
| SMART | SM00355 | 11 | 1097 | 1119 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.1E-5 | 1120 | 1148 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0037 | 1120 | 1144 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.466 | 1120 | 1149 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1122 | 1144 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.62E-9 | 1136 | 1178 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.5E-8 | 1149 | 1174 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0014 | 1150 | 1174 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.741 | 1150 | 1179 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1152 | 1174 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.71E-8 | 1168 | 1202 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 2.9E-9 | 1175 | 1203 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.79 | 1180 | 1206 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.889 | 1180 | 1211 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 1182 | 1206 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1213 aa Download sequence Send to blast |
MLTVGETNWN MRGVSRSKGS LLRFMKEEIP GVTSPMVYVA MMFSWFAWHV EDHDLHSLNY 60 LHMGAGKTWY GVPRDAAVAF EEVVRVQGYG GEVNPLITFA VLGEKTTVMS PEVLVGAGVP 120 CCRLVQNAGE FVVTFPRAYH SGFSHGFNCG EAANIATPEW LLFAKEAAIR RASINYPPMV 180 SHFQLLYDLA LAFCSRVPTC INTGPRSSRL KDKKKGEGES MVKEQFVQNC IENNKLLHIL 240 GKDSTVVRLP ESSSDISVCS NLRVGSHLRV NRSLSEDTMM SHGGLVSDNL IADRKQTIDL 300 VDDLYSMKAF SSFYERKRIL SMYGAGDIHP PRDGEGDSKA QGDRLSDQRL FSCVTCGILT 360 FACAAIVQPR EPAARYLMSA DCSFFNDWTV NPGVPTKGFN VSAGADIIAY KRRTQAGSVE 420 KSRAAALYDV SPLQSTNYPI QEAHQSSEAF SNSDKLKGTS ALGLLALNYG NSSDSEEDQI 480 DSDAPNETTS RPQYGNNNLP DSALPPFLPE HHSGPNGGSP HNCCPEIGYR GANFVDKSHQ 540 TFKYSANFRA DETRSSEKHG STGFPKSDED SSRMHIFCLE HAIEVDQRLR PIGGVYIYLA 600 CHPDYPRVEA EAKLLAEELG IDHSWNEIAF RDAMGEDKER IQSALDSEEA IAGYGDWAVK 660 LGINLFYSAN LCSSPLYSKQ MPYNSVIYNA FGRESSSSPA KLDDFGRRPS KQKKSVAGKW 720 CGKVWMSNQV HPFLAPKDAE EVEEERSFHA SMTSDDKLER QSGLNRETTL ATRKFSRKRK 780 MARESGPSKK KSVSRKEEVS DDALAEDSEK LMSIPKHKTA KSFRREDPVS DDQVDEISCL 840 QHQTIPRRKQ KKFVKRGYAS SDDDSLEDKF KALRGKNFRK ATNFTSSGDM LSDDSLEEDS 900 QQQGRILTGK KTKHYERDDV MSDDSEGNYN KLQHPRIRKK TPANAKFEGA DSDSSPEDAF 960 PQRQRSLRNR KVNRDTLRIK NQGNLLEKLK KRGIPRVTKE VSARHLKQES SRPRSNKYER 1020 SVEQSDESDE EDQEGGGPST RLRKRTAKPT KERKADKPPP GKKQANNGKK NVKAGPTVKR 1080 PVGRNDARME KFEEAEYKCD LEGCTMSFGS KQELTLHKKN VCPVKGCGKK FFSHKYLVQH 1140 RRVHLDDRPL KCPWKGCKMT FKWAWARTEH IRVHTGARPY ICAEPGCGQT FRFVSDFSRH 1200 KRKTGHSVKG KG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6a57_A | 3e-68 | 1095 | 1209 | 21 | 135 | Lysine-specific demethylase REF6 |
| 6a58_A | 3e-68 | 1095 | 1209 | 21 | 135 | Lysine-specific demethylase REF6 |
| 6a59_A | 3e-68 | 1095 | 1209 | 21 | 135 | Lysine-specific demethylase REF6 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 775 | 780 | SRKRKM |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
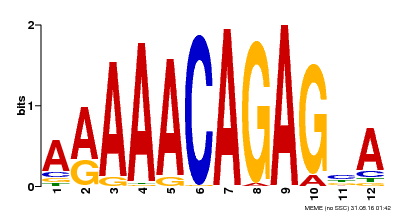
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Eucgr.K02763.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010037473.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | A0A059A5S0 | 0.0 | A0A059A5S0_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010037473.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5259 | 28 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Eucgr.K02763.1.p |



