 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Eucgr.H04783.1.p | ||||||||
| Common Name | EUGRSUZ_H04783, LOC104415786 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Myrtales; Myrtaceae; Myrtoideae; Eucalypteae; Eucalyptus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1048aa MW: 117034 Da PI: 5.4111 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 179.1 | 5.5e-56 | 20 | 136 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptf 94
l+ ++rwl++ ei++iL+n++k++++ e+++ p+sgsl+L++rk++ryfrkDG++w+kk+dgktv+E+he+LK g+++vl+cyYah+e+n++f
Eucgr.H04783.1.p 20 LDAQHRWLRPAEICEILQNYKKFQIAPEPANMPPSGSLFLFDRKVLRYFRKDGHNWRKKRDGKTVKEAHERLKAGSIDVLHCYYAHGEHNENF 112
6779***************************************************************************************** PP
CG-1 95 qrrcywlLeeelekivlvhylevk 118
qrr+yw+Leeel++ivlvhylevk
Eucgr.H04783.1.p 113 QRRTYWMLEEELSHIVLVHYLEVK 136
*********************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 80.455 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.7E-77 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 3.3E-49 | 22 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF01833 | 2.2E-4 | 464 | 544 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 8.92E-16 | 464 | 550 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:2.60.40.10 | 6.3E-4 | 464 | 539 | IPR013783 | Immunoglobulin-like fold |
| CDD | cd00204 | 1.23E-13 | 640 | 755 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 3.4E-17 | 641 | 758 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 17.873 | 644 | 757 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 9.17E-17 | 644 | 757 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 8.2E-8 | 668 | 759 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.671 | 696 | 728 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.011 | 696 | 725 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 190 | 735 | 764 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 1.78E-7 | 866 | 921 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.13 | 870 | 892 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.48 | 871 | 900 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0042 | 872 | 891 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.36 | 893 | 915 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.956 | 894 | 918 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0011 | 896 | 913 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1048 aa Download sequence Send to blast |
MADARRYVLG NPLDIQQILL DAQHRWLRPA EICEILQNYK KFQIAPEPAN MPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKRD GKTVKEAHER LKAGSIDVLH CYYAHGEHNE NFQRRTYWML 120 EEELSHIVLV HYLEVKGNRM NIHRIKDTEE TISLSHETEV NHETNSSLSS ISHQNNYQVP 180 SQTTDSSSPQ ASEYEDAESA YSNQANSGNH SIVDFQEAGM QNIDTGHSDP YYPASDSTDD 240 YQQNLSLKPG MDFLSLSRAD ITRECNVAGL ALEPQKNLDF SSWEPLLESC TSGGQSTLIQ 300 VLGSSAQPVT PDIMPKHAHD TVEQLFTDGI STKQEIGIQS QDQEELQVHL TNKIAESVST 360 ENGHHSNSFL DGNVALEVKA GCPSMGKQPF ANILAEEGLK KMDSFNRWMS KQLGDVNALL 420 GQSSTGAYWS AIVNDDGIDS SDSQEHDNNF MLTPSLSQEQ LFSIIDFSPS WTYADLEIKV 480 LITGRFLRSQ QDAELCKWSC MFGEVEVPAE VMADGVLRCH APVHKAGRVP FYVTCSNRVA 540 CSELREFEFR VNNSQDVISA DTKDGSINET LYMRFTKLLC SNTPRLSNSN IGENSQLRSK 600 ICSLLKDDDD AHWDHVMEST SDEFSTEGVR EQFFQKLLKE KLCEWLIEKA AEGGKGASVL 660 DEDGQGVLHF ASALGYDWAL QPTIIAGVSV NFRDANGWTA LHWAASFGRE RTVASLISLG 720 AAPGALTDPT PKYTTGRTPA DLASANGHKG IAGYLAESAL SAHLSLLNLD TAGGDASHGS 780 GGKAVQTISE RSPTPIREGD LPSGVSLKDS LAAVCNATQA AARIHQVFRV QSFQRKQLRE 840 FGDDNHGISD EHALTLVASK VYKPGQVDEP LHAAAVRIQN KFRGWKGRKE FLITRQRIVK 900 IQAHFRGHQV RKNRKNILWS VGIVEKVILR WRRKGSGLRG FKREELTDTS SMQDASSDED 960 EYDFLKEGRK QTEERLEKAL ARVKSMVQYP EARDQYRRLL NVVSEIQEAQ VEYEMTPDHN 1020 GEATDFDGDL TDLEQLLDDD TYMHLGP* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
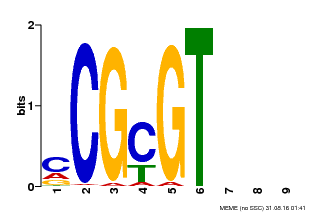
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Eucgr.H04783.1.p |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010025463.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 3 isoform X1 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A059B7C0 | 0.0 | A0A059B7C0_EUCGR; Uncharacterized protein | ||||
| STRING | XP_010025463.1 | 0.0 | (Eucalyptus grandis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM8601 | 25 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Eucgr.H04783.1.p |
| Entrez Gene | 104415786 |



