 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Dusal.0157s00007.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Chlorophyceae; Chlamydomonadales; Dunaliellaceae; Dunaliella
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 167aa MW: 18970.4 Da PI: 11.0962 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 56 | 9.7e-18 | 61 | 110 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrd.psengkr.krfslgkfgtaeeAakaaiaarkkleg 55
gy+GVr++ +g ++AeIrd + kr ++g+fgtaeeAa a+++a+ +l+g
Dusal.0157s00007.1.p 61 GYRGVRQRA-WGTYAAEIRDaK-----HkKRRWIGTFGTAEEAACAYDQAAVELHG 110
8*****998.**********44.....45*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00380 | 1.3E-26 | 61 | 124 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.994 | 61 | 118 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 7.5E-11 | 61 | 110 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.9E-19 | 61 | 120 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 3.7E-25 | 62 | 118 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.51E-23 | 62 | 120 | No hit | No description |
| PRINTS | PR00367 | 3.5E-8 | 62 | 73 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 3.5E-8 | 84 | 100 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0042991 | Biological Process | transcription factor import into nucleus | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 167 aa Download sequence Send to blast |
MVFPPLVSSP NFRMSATSSQ SSSGSSISAF AEDISAVRDQ LRVDVPERRF PEEKKHKNRH 60 GYRGVRQRAW GTYAAEIRDA KHKKRRWIGT FGTAEEAACA YDQAAVELHG LHAHTNFEWS 120 KEPDIRTSAF QQQKGRKKRK ALQVRINGPR VFFKALNVFK QTPSHF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 1e-16 | 60 | 117 | 12 | 71 | Ethylene-responsive transcription factor ERF096 |
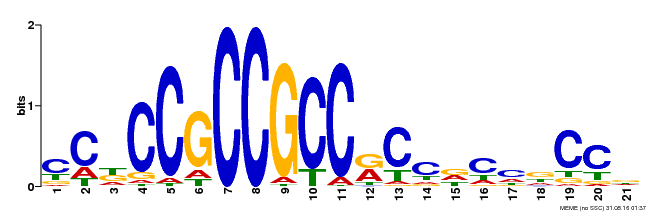
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00456 | DAP | Transfer from AT4G27950 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027362376.1 | 5e-22 | ethylene-responsive transcription factor ESR2-like | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP132 | 15 | 40 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G27950.1 | 2e-14 | cytokinin response factor 4 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Dusal.0157s00007.1.p |



