 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Dusal.0059s00028.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Chlorophyceae; Chlamydomonadales; Dunaliellaceae; Dunaliella
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 1116aa MW: 115986 Da PI: 6.4325 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 41.1 | 4.1e-13 | 35 | 79 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +W++eE+ ++++a +++ + Wk+I +++g ++++ q++s+ qk+
Dusal.0059s00028.1.p 35 RESWSEEEHGRFLEALRLYARD-WKRIEEHVG-TKSATQIRSHAQKH 79
789****************977.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.4E-13 | 30 | 82 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 7.9E-7 | 32 | 88 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 16.103 | 32 | 84 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 3.5E-13 | 33 | 82 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 3.0E-10 | 34 | 82 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.0E-11 | 35 | 79 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 9.07E-8 | 37 | 80 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0032922 | Biological Process | circadian regulation of gene expression | ||||
| GO:0043966 | Biological Process | histone H3 acetylation | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1116 aa Download sequence Send to blast |
MREERAEESG PDQRPGQAGT VPPKPRKAYV LTKTRESWSE EEHGRFLEAL RLYARDWKRI 60 EEHVGTKSAT QIRSHAQKHF HKAMQQGEEI PPPRAKRRSA APTPRSRPLP KQQQQVARAP 120 QRQPSHRQQQ QQQQQQQDDY QQASHAWLRR GKNGSGASQF LEHRGQHQGS GSTGQDARHL 180 GVTIEQDSSG IAELMCSMNP AMMGQDAGDA GLLSSLGMLQ PTLLGCPASL MGQVGPGLPL 240 GGYFPGTGGF LGGDSSMITW AQQPQQQQQQ QQQLTDFVGG ASFSSTIAAS QAVLEAMDGA 300 GTGGSGCMPL GELPQQQVPC KAMCGSDAAG AAGGPLEGTV GGSSSSSNLA AKQVAQQQQQ 360 QQQQQGGSSS LAAAAAATTG GLLSGHLAIL PGGTRVEGGM GAAEFSQLMS MTAIATAQGQ 420 VQGQVQHCQE HGQQLMRVGD QDEAFVVGRQ SCEGGMDRQQ GTASSEEDQS IGSNQQGSGT 480 DGDGCGSMKD GSRTNGQGHS EVAKANQEPG TWAGTQQQPQ QNLPGPVHSS GRSVREGNGS 540 GATTTTTAGC GTKRLHSVDA AAASTDDEDS LCLADVVLRP HPKGRRTSHT SRQTASSAAG 600 VAPGPSPSPA PASPPHHKHN SPASLPPFEA SQPITLLGRA SVAPNSSSSS FSHRVRARSS 660 SAQSHSHAHP VHHQREGTPV ANTHRLARMQ QRQEQEQEPE QKVQLMGAHQ GPHPSAGAGW 720 VGDSSRSSMD HRHLTSFQNS QGNSGLRATA AHHAPQGVKV EEGCSGGEHN SRSVPQTDFE 780 WMIREFEQID DDSGPCCGYN WMSTGPTSGV EQQQQQQVAE RQQQLLLMQQ REQHLSSYLR 840 GTTPPHASSM GSGRTPGHGS GAGAQQELFL QHGGGAGAQH VPGSVCMEGL QAAYPKQLEQ 900 HQQQQQQQQL GVVVAAAAGL AHAAGGDSGE GDVGSNSTDD SDPGPGTHSE CEGRVPSSHA 960 GMCPGQPLPN GLSPALFDGP GRVPTPLPFD MAPTPDLPCS ISPSPQLALA ADGLPLPLEP 1020 LLTFGPRSSP PDSATLMPLD TPSPPVVSND SALGAGGGGV VGAGNCGGHG GESENEVGNG 1080 PQALRHALSL PEGYGPAVTC TQSSGSMADI DALLN* |
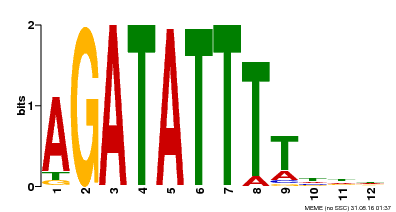
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00338 | DAP | Transfer from AT3G09600 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP454 | 14 | 27 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G09600.2 | 5e-24 | MYB_related family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Dusal.0059s00028.1.p |



