 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Do016252.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Dichantheliinae; Dichanthelium
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1000aa MW: 111113 Da PI: 5.597 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 180.5 | 2e-56 | 21 | 136 | 2 | 117 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcyw 100
l+ ++rwl+++ei++iL n++k++++ e+++rp+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg+v+vl+cyYah+een++fqrr+yw
Do016252.1 21 LEAQNRWLRPTEICQILSNYKKFSIAPEPPNRPPSGSLFLFDRKILRYFRKDGHNWRKKKDGKTVKEAHEKLKVGSVDVLHCYYAHGEENENFQRRTYW 119
5679*********************************************************************************************** PP
CG-1 101 lLeeelekivlvhylev 117
lLee + +ivlvhyle
Do016252.1 120 LLEEGFMNIVLVHYLEI 136
**************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 80.736 | 16 | 142 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 4.2E-76 | 19 | 137 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 1.1E-50 | 22 | 135 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 3.5E-6 | 403 | 490 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 1.1E-5 | 404 | 477 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 6.47E-18 | 404 | 490 | IPR014756 | Immunoglobulin E-set |
| SuperFamily | SSF48403 | 2.02E-17 | 581 | 696 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.54E-14 | 582 | 694 | No hit | No description |
| PROSITE profile | PS50297 | 16.945 | 585 | 696 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 7.2E-18 | 586 | 697 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 1.8E-7 | 608 | 697 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.028 | 635 | 664 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.897 | 635 | 667 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 460 | 674 | 703 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 6.37E-8 | 809 | 860 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.14 | 809 | 831 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.682 | 811 | 839 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0054 | 811 | 830 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.012 | 832 | 854 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.633 | 833 | 857 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.2E-4 | 835 | 854 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0050826 | Biological Process | response to freezing | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1000 aa Download sequence Send to blast |
MAEIRKYAMS NQPPDIPQIL LEAQNRWLRP TEICQILSNY KKFSIAPEPP NRPPSGSLFL 60 FDRKILRYFR KDGHNWRKKK DGKTVKEAHE KLKVGSVDVL HCYYAHGEEN ENFQRRTYWL 120 LEEGFMNIVL VHYLEIKGGK QNFNRAKEAE ENAGLSNADS PACSNSFASQ SQVASQSMDA 180 ESPISGQISE YEDAETDAGY LGEMQPTPAN FTNNFVSHND ISSAFNETGA GLRGNPKTPI 240 DSVRFGESFP EYPGGFTEPT LYSSVATVGN NLDDSLQTFM SEALYTNNLT QKELDALSAA 300 GITSSQAEND GYTDQSARYP LLKQSSLDLF KIESDGLKKF DSFSRWMSSE LAEVADLDIK 360 SSSDAFWSTT ETVNVADGSG LPINEQLDAF VVSPSLSQDQ LFSIIDVSPS WAYNGTKTKV 420 LITGTFLAKK EDVENCRWSC MFGDVEVSAE VLVDGTLRCY TPVHHSGRVP FYVTCSNRVA 480 CSEVREFEFR DSETHYMETS DLHPTGINEM HLHIRLDKLL SLGPEDYEKY VLSNGNKAEL 540 IDTINSLMLD DKLSNLALSS DEKELSTVRD QNIEKQVKEK LYYWLIHKIY DDGKGPNVLG 600 KEGQGVIHLV AALGYDWAIK PIVAAGVNVN FRDIRGWTAL HWAASCGRER TVCALIANGA 660 AAGYLTDPTQ QFPSGRTAAD LASENGHKGI AGFLAESALT SHLSALTLKE SQGGKAEEIC 720 GLAAAEDFAE PSSVQPASVD SQAESLKDSL GAVRKSTQAA ARIFQAFRVE SFHRKKVIEY 780 GDDDCGLSDE RTLSLVSLKN AKPGHSDMPV HSAAVRIQNK FRGWKGRKEF MIIRQKIVKI 840 QAHIRGHQVR KSYRKVVWSV GIVEKVILRW RRKGRGLRGF QAEKQLEGPS SQIQPAKSEA 900 EDEYDFLKDG RKQAEGRLQR ALARVHSMTQ YPEARDQYRR LQTCVNELQE PQVLSLFSTI 960 IVAMQDRVLS DSAGADGGDL MAELEELCRG DGDAPMSTIS |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00501 | DAP | Transfer from AT5G09410 | Download |
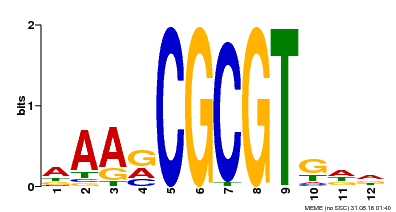
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Do016252.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025800735.1 | 0.0 | calmodulin-binding transcription activator 3-like | ||||
| TrEMBL | A0A1E5VCX2 | 0.0 | A0A1E5VCX2_9POAL; Calmodulin-binding transcription activator 1 | ||||
| STRING | Pavir.Ba00386.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.1 | 0.0 | ethylene induced calmodulin binding protein | ||||



