 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Do008052.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Panicoideae; Panicodae; Paniceae; Dichantheliinae; Dichanthelium
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1157aa MW: 129068 Da PI: 6.2829 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 176.4 | 3.6e-55 | 19 | 136 | 2 | 118 |
CG-1 2 lke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcy 99
lke ++rwl++ ei++iL+n+++++++ e+++rp+sgsl+L++rk++ryfrkDG++w+kk+dgkt++E+he+LK g+++vl+cyYah+een +fqrr+y
Do008052.1 19 LKEaQHRWLRPAEICEILKNYRNFHIAPEPPNRPPSGSLFLFDRKVLRYFRKDGHNWRKKRDGKTIKEAHERLKSGSIDVLHCYYAHGEENGNFQRRSY 117
45559********************************************************************************************** PP
CG-1 100 wlLeeelekivlvhylevk 118
w+Lee++ +ivlvhylevk
Do008052.1 118 WMLEEDFMHIVLVHYLEVK 136
****************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.321 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.0E-77 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 8.1E-49 | 21 | 135 | IPR005559 | CG-1 DNA-binding domain |
| CDD | cd00102 | 3.06E-4 | 424 | 511 | No hit | No description |
| Gene3D | G3DSA:2.60.40.10 | 5.3E-6 | 425 | 509 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 2.62E-17 | 425 | 510 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 5.3E-8 | 428 | 504 | IPR002909 | IPT domain |
| SuperFamily | SSF48403 | 1.18E-15 | 616 | 720 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 7.13E-12 | 619 | 717 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 7.1E-16 | 619 | 723 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.923 | 625 | 657 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50297 | 17.343 | 625 | 729 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.603 | 658 | 690 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 3.19E-8 | 830 | 882 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 0.075 | 831 | 853 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.407 | 832 | 861 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0038 | 833 | 852 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0022 | 854 | 876 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.56 | 855 | 879 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.3E-4 | 857 | 876 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1157 aa Download sequence Send to blast |
MAEARRYAIA PQLDIEQILK EAQHRWLRPA EICEILKNYR NFHIAPEPPN RPPSGSLFLF 60 DRKVLRYFRK DGHNWRKKRD GKTIKEAHER LKSGSIDVLH CYYAHGEENG NFQRRSYWML 120 EEDFMHIVLV HYLEVKGGKS SSRIRGHDDM LQAARTDSPL SQLPSQTTQG ESSLSGQASE 180 YEETESVYCH YGPKLSWPVF FCYPYLGNHQ GFPAIATTTD FYSHGQDALP VALNEPGLGI 240 TFNGADNQLD PSLLNGLVKP DQGVHIMPPP QITDPSKQFP FTEGPGIESF TFDEVYSNGL 300 SIKDADTIGT DEESLWQLPG AISSFPTEDG LQQNYRSLDE AINTLLKTQS SSLSDILKDS 360 FKKSDSFTRW MSKELCEVDD SQIKSSSGVY WSSEETGNSI EASSRDQLDQ FTVDPVLAQD 420 QLFSIFVFSP SWTYAGSKTR VLITGRFLNS DEVQRCKWSC MFGEVEVPAE ISGDGTLRCY 480 SPLHKPGRVP FYVTCSNRLA CSEIREFEFR PSDSQHMDGP GPHDAANKTY LQMRLDDLLS 540 LGQDEYQAMV SNPTKEMTDL SKKISSLMTD NDSWSELLKL AGDNELATDD KQDQFFEKRL 600 KEKLHTWLVH KTGDGGKGPS VLDEEGHGVL HLAAALGYDW AIRPTISAGV NINFRDAHGW 660 TALHWAAFCG RERTVVALIA LGAASGALTD PTPDFPTGST PADLASANGY KGISGFLAES 720 SLTSHLQTLN LKEAMGSHAS EISGLPGIGD VTERRASPLA GEGLLAGSMG DSLGAVRNAS 780 QAAARIYQVF RMQSFQRKQA VQYEDDNGAI SDDRALSLLS VKPSKPGQLD PLHAAATRIQ 840 NKFRGWKGRK EFLLIRQRIV KIQAHVRGHQ VRKHYRKIVW SVGIVEKVIL RWRRRGAGLR 900 GFRSTEGAVE GTSRSGSDVI QNKPAEDDYD FLQRGRKQTE ERLQKALARV KSMVQYPDAR 960 DQYKRILTAV TRLQESQAMQ EKMLESSTDM DEGFVMTEFK ELWDDDMPMP GFPPSRSPSP 1020 GRFEPLPPSP YAAAAMDAAP SAFMVLTGHL VLDELAVRPP KKWAKIQCAS KKAHGCGKHG 1080 EELVEDLSLW LSLFDGEDDL NSSLCIRMSD DALRSVQADL GISRSMVDAE GSVQIANQER 1140 LVLTLFLRRQ CCVKRAY |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
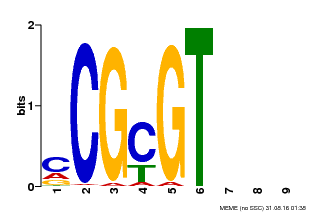
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Do008052.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004985392.1 | 0.0 | calmodulin-binding transcription activator 3 isoform X1 | ||||
| TrEMBL | A0A1E5W710 | 0.0 | A0A1E5W710_9POAL; Calmodulin-binding transcription activator 3 | ||||
| STRING | Si034076m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



