 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV53764.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 426aa MW: 48109.6 Da PI: 5.1583 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.8 | 1.8e-18 | 49 | 102 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k++++t +q+++Le+ Fe ++++ e++ +LAk++gL+ rqV vWFqNrRa++k
KZV53764.1 49 KKRRLTVDQVKALEKSFEVENKLEPERKVKLAKEIGLQPRQVAVWFQNRRARYK 102
56689************************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 127.1 | 7.7e-41 | 48 | 140 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekkrrl+ +qvk+LE+sFe e+kLeperKv+la+e+glqprqvavWFqnrRAR+ktkqlE+dy Lk+++d+lk++ ++L++++++L ++++e
KZV53764.1 48 EKKRRLTVDQVKALEKSFEVENKLEPERKVKLAKEIGLQPRQVAVWFQNRRARYKTKQLERDYGILKANFDNLKQNYDTLHSDNQALLRQIQE 140
69*************************************************************************************999986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.18E-18 | 41 | 106 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.313 | 44 | 104 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 6.2E-18 | 47 | 108 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.74E-16 | 49 | 105 | No hit | No description |
| Pfam | PF00046 | 1.2E-15 | 49 | 102 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-19 | 51 | 110 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 3.6E-5 | 75 | 84 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 79 | 102 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 3.6E-5 | 84 | 100 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 6.6E-17 | 104 | 145 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 426 aa Download sequence Send to blast |
MKRLGSSDSL SALMSTDELN QESEPIYSRE FRTMLEGLDE EEPGHVSEKK RRLTVDQVKA 60 LEKSFEVENK LEPERKVKLA KEIGLQPRQV AVWFQNRRAR YKTKQLERDY GILKANFDNL 120 KQNYDTLHSD NQALLRQIQE LKSKVNGDDV KSERRAISVK EEAMVLSDTE EKNNLETPWN 180 DNVSSFLDFK DGSSDSTDSS AILNEGNNSP HQFTGSSVVG DETKAVKATS LIGDDSKFVE 240 YNFLGEDHDQ SCSSLFSDEQ APTLHCTLQA RPVSSSGTSP EDHVPTTEEE SILDMPFLIQ 300 AYPVRIHFPY LKILGRYNFP YLVDEIVENR GRSPRRSWRL YRPSEISAEI CPSQPPSPPI 360 NTRFAPSFWT SHSDSHTLAL TAHSPLSFLS GTLPDLGIGG ASSDMLPAPL TRDLWLQVPR 420 RSAFRL |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 96 | 104 | RRARYKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may function as a negative regulator of the flowering time response to photoperiod. May act to repress cell expansion during plant development. {ECO:0000269|PubMed:14623244}. | |||||
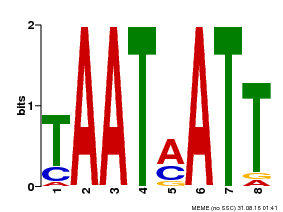
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00051 | PBM | Transfer from AT4G40060 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV53764.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022884552.1 | 8e-84 | homeobox-leucine zipper protein ATHB-6-like isoform X1 | ||||
| Swissprot | Q940J1 | 4e-64 | ATB16_ARATH; Homeobox-leucine zipper protein ATHB-16 | ||||
| TrEMBL | A0A2G9I4D6 | 6e-90 | A0A2G9I4D6_9LAMI; Transcription factor HEX | ||||
| STRING | Migut.L00161.1.p | 7e-76 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2718 | 24 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G40060.1 | 2e-66 | homeobox protein 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



