 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | KZV30100.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Gesneriaceae; Didymocarpoideae; Trichosporeae; Loxocarpinae; Dorcoceras
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 294aa MW: 33263.5 Da PI: 4.7768 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.4 | 2.5e-18 | 55 | 106 | 5 | 56 |
SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 5 ttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ eq+++Le+ Fe ++++ +++ +LA++lgL+ rqV +WFqNrRa++k
KZV30100.1 55 RRLGVEQVKALEKNFEVENKLEPDRKVKLAQELGLQPRQVAIWFQNRRARWK 106
467789*********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 130.1 | 8.9e-42 | 52 | 144 | 1 | 93 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekeveeLreelke 93
ekkrrl eqvk+LE++Fe e+kLep+rKv+la+eLglqprqva+WFqnrRAR+ktkqlE+dy+ Lk++ydalk++ e+L++++e L++e++e
KZV30100.1 52 EKKRRLGVEQVKALEKNFEVENKLEPDRKVKLAQELGLQPRQVAIWFQNRRARWKTKQLERDYDILKASYDALKHSYENLQRDNEILTKEIQE 144
69***************************************************************************************9987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 6.84E-19 | 44 | 110 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 16.973 | 48 | 108 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.8E-17 | 51 | 112 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.48E-16 | 53 | 109 | No hit | No description |
| Pfam | PF00046 | 1.7E-15 | 54 | 106 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-19 | 57 | 115 | IPR009057 | Homeodomain-like |
| PRINTS | PR00031 | 7.0E-6 | 79 | 88 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 83 | 106 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 7.0E-6 | 88 | 104 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 2.7E-17 | 108 | 148 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0009788 | Biological Process | negative regulation of abscisic acid-activated signaling pathway | ||||
| GO:0030308 | Biological Process | negative regulation of cell growth | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048510 | Biological Process | regulation of timing of transition from vegetative to reproductive phase | ||||
| GO:0048573 | Biological Process | photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 294 aa Download sequence Send to blast |
MKRPNSSDSL GDLMSICPES TDEHDQENNP IYSGEFGTFV EGLVEESKHV MEKKRRLGVE 60 QVKALEKNFE VENKLEPDRK VKLAQELGLQ PRQVAIWFQN RRARWKTKQL ERDYDILKAS 120 YDALKHSYEN LQRDNEILTK EIQELKSKDG EDGKSESKVS AKEEATMVSD VEKGKLGHET 180 VACEHDNNHV SYIGNNGSSD STDSSAILNE DNSPHQFLDS KNGSGSTSSM NLSFKFSDFK 240 VSAASVVADA HNPHTQLQSV KIEEHNFLDE ESCSSLFSDA QAPTLHWYFS DEWN |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 100 | 108 | RRARWKTKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00271 | DAP | Transfer from AT2G22430 | Download |
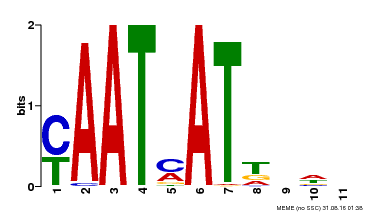
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | KZV30100.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011097532.1 | 1e-106 | homeobox-leucine zipper protein ATHB-6 | ||||
| Refseq | XP_022884552.1 | 1e-106 | homeobox-leucine zipper protein ATHB-6-like isoform X1 | ||||
| TrEMBL | A0A2Z7B8R5 | 0.0 | A0A2Z7B8R5_9LAMI; Homeobox-leucine zipper protein ATHB-6-like | ||||
| STRING | Migut.H02327.1.p | 4e-92 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2718 | 24 | 53 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22430.1 | 8e-62 | homeobox protein 6 | ||||



