 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Dca46872.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Caryophyllaceae; Caryophylleae; Dianthus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1449aa MW: 160369 Da PI: 7.9809 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 15.9 | 3.6e-05 | 1307 | 1329 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp dCgk+F ++ +L++H r+H
Dca46872.1 1307 CPvkDCGKTFFSHKYLVQHRRVH 1329
9999*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 1.9E-15 | 26 | 67 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 13.578 | 27 | 68 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 2.2E-13 | 28 | 61 | IPR003349 | JmjN domain |
| SMART | SM00558 | 1.8E-50 | 205 | 374 | IPR003347 | JmjC domain |
| PROSITE profile | PS51184 | 33.308 | 205 | 374 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 2.33E-25 | 223 | 388 | No hit | No description |
| Pfam | PF02373 | 1.0E-37 | 238 | 357 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 8.6E-4 | 1275 | 1304 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 10 | 1282 | 1304 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 13.11 | 1305 | 1334 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0045 | 1305 | 1329 | IPR015880 | Zinc finger, C2H2-like |
| SuperFamily | SSF57667 | 1.18E-6 | 1305 | 1341 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.7E-5 | 1306 | 1334 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1307 | 1329 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1449 aa Download sequence Send to blast |
MGSGEHSSSS SSSSTEVFPW LKTLPVAPEY HPTIQEFQDP ISYIFKIEPE ASKFGICKII 60 PPVPPPSKKS IVSNFNKSIS LLKNSSSSGG GGGAAEFTTR QQQVGFCPRK QRRPVQKSVW 120 RSGESYTFQQ FEAKARNFEK NYIKNNVKSG GSCKKTVLTP MEVETLYWRA NGGRPFSVEY 180 ANDMPGSGFG GVRERKRGAG DVVSNVGESG WNMRGVARSN MSLLRFMKED IPGVTSPMVY 240 VAMMFSWFAW HAEDHDLHSL NYLHFGSPKT WYGVPRDAAF AFEDVIRVHG FGDEINPLVT 300 FATLGEKTTV MSPEVLVNAG VPCCRLVQNA GEFVITFPRA YHSGFSHGFN CGEASNIATP 360 EWLRFAKDAA IRRAAINYPP MVSHYQLLYD LALAIRVPAG TTSEPRSSRL KDKKKGEGEM 420 LVKQMFVEDV MHNNELLYTL GHGSEVVLLP HNSSEKFVWS NLRVGSKYKV KPGLPFSFHS 480 SEDPAKAYDD IMLARDIKQH KAFYSVKAKL NSMSHKTQQS ETESKEDAAA DGLSDRGLFS 540 CVKCGIWTFA CVAIVQPTEK AAQYLMSADC SVFKDWMAGS HAINGDATAS QPESTSGFMK 600 KNAFGGFYDV HIQSADCRVQ STEGSPIIKS DTAKPREESS ALGLLALQYG NTTDSEEDDD 660 ALPSHPAISD NNLSEGGSSD AGRHQDNATS PNSEQGYASG LEKGPTVNSS RSECDDEHSP 720 HRSNSYERSD HGRASSADSE NATHNFSGKF TEEKNFVKSS GGPELVTEPK LPYTGCDEDS 780 SRMHVFCLEH AVEVEKLLQP IGGVHVVLLC HPDYPNVQFE AMKVAEELEV NYVWKDITFT 840 EATKVDEERI QMALQSEEAT PKTGDWSVKL GINLFYSAIL SRSPLYSKQM PYNSVIYDVF 900 GCGSPSKSSP DAKSGQKGYS RQRKIVVAGK WCGKVWMSNQ VHPLLLQSDP PEKEKVLGLC 960 STTAENPRRK PETGNKPQTT SRNRKPTKRK LSVTSRVAKK MKFELNEVAD SDPEDSPNDR 1020 IESAPRRSIR KLKTQRRSIR RSPDSLSDER IEPLSRISIR KPKTQQRLVK SATHHEVESE 1080 DGVSEDSSGH DEPSQVPLRG PRVLKGGRIS VPKLRVQQNV SLDEPSDDSA DDDVSHHHVL 1140 RRCSSKTKAA RKPTRQLKCS DSEGSLSDES REEHTVVGSP EFEPEPTSED ELRGGPSTRL 1200 RKRISKPSAR VLETKEAARS LAKNSRGKKI SVVVRNPPER KGPKMRLPAM KGPKMRLPAK 1260 KAPAVSSDED TENESDDGEG EFKCDIEGCT MSFQSKQALA LHKRNICPVK DCGKTFFSHK 1320 YLVQHRRVHV DDRPLKCPWK GLNASVYLRC MSELVVEDVV LAPSSLNYGE LESDLLTLVI 1380 AVNVSVYLRC TWELVVEDVA LAPSSLNYGY CTLVRHGGNS VTHFVIRSTT FVGDASVCDF 1440 PLVLLKIAF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 4e-76 | 13 | 395 | 1 | 353 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 4e-76 | 13 | 395 | 1 | 353 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 983 | 990 | RKPTKRKL |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
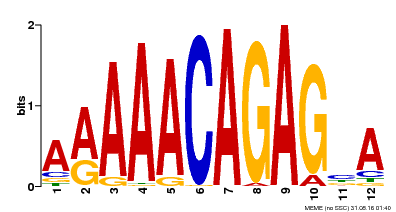
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010667395.1 | 0.0 | PREDICTED: lysine-specific demethylase REF6 isoform X1 | ||||
| TrEMBL | A0A0J8B695 | 0.0 | A0A0J8B695_BETVU; Uncharacterized protein | ||||
| STRING | XP_010667395.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||



