 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Dca41449.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Caryophyllaceae; Caryophylleae; Dianthus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 591aa MW: 65606.7 Da PI: 6.9338 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 40.2 | 5.9e-13 | 434 | 480 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ +K++Ka+ L A+ YI +Lq
Dca41449.1 434 NHVEAERQRREKLNQRFYALRAVVPNI-----TKMDKASLLGDAISYITELQ 480
799***********************7.....7*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 3.4E-48 | 49 | 236 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.984 | 430 | 479 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 9.48E-15 | 433 | 484 | No hit | No description |
| SuperFamily | SSF47459 | 7.46E-18 | 433 | 491 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.6E-17 | 434 | 493 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.7E-10 | 434 | 480 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.6E-17 | 436 | 485 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 591 aa Download sequence Send to blast |
MGAKSMVLNE EDKAMIASIL GTRAFDYLID RPYSAESSLV SAEISENIQN KLVDLVEKPN 60 TDNFSWNYGF FWQISRSRGG ELVLSWGDGY CREPIDGEES EATRFLKYRV EDEGQQKIRK 120 RVLQKLNGFF GGSDDDNLAL RLDRVTDMEM FFLLSMYFSF PRCEGGPGKC FASGKHVWEA 180 DLLGSELDYC VRSFLAKSAG IQTVVLIPTE FGVVELGSVR KVSENHEMLS ALKCAFAFSN 240 NTTTPTTLPL LTPKVEEDAS TAPGFANLRF VNGVDGFPKI FGQHLSATVR PQFREKLAVR 300 KTEDKSWEPN VNGNKLCFTT QTPRNGLQSP PSWSNISGVK HATSMEVYSP QNSRLQDFAN 360 GIRGDCRSNN NQSQQKKPVQ MQIDFTGATS RAGSADSEHS DPKPAYGTEY GPSDDSKTRK 420 RGRKPANGRE GALNHVEAER QRREKLNQRF YALRAVVPNI TKMDKASLLG DAISYITELQ 480 KKLKDLESNS NGDAQRLSQV PNSDVEVQAA DGEVIVRVRC PLDSHPVAPV MQAFKNEAIQ 540 VLNSKLASDT DTVSHTFVIK SQSSEQLTKD KLLAMFSGES KPLQTRSPTG E |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 2e-25 | 428 | 500 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_B | 2e-25 | 428 | 500 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_E | 2e-25 | 428 | 500 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_F | 2e-25 | 428 | 500 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_G | 2e-25 | 428 | 500 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_I | 2e-25 | 428 | 500 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_M | 2e-25 | 428 | 500 | 2 | 74 | Transcription factor MYC2 |
| 5gnj_N | 2e-25 | 428 | 500 | 2 | 74 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
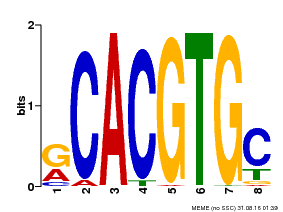
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010696336.1 | 0.0 | PREDICTED: transcription factor bHLH13 | ||||
| Swissprot | A0A3Q7ELQ2 | 0.0 | MTB1_SOLLC; Transcription factor MTB1 | ||||
| TrEMBL | A0A167V6C1 | 0.0 | A0A167V6C1_CAMSI; MYC2 trancriptor | ||||
| STRING | XP_010696336.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G01260.3 | 1e-175 | bHLH family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



