 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Dca27529.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; Caryophyllales; Caryophyllaceae; Caryophylleae; Dianthus
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 648aa MW: 70893.6 Da PI: 6.6824 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 91.4 | 7.7e-29 | 211 | 264 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQkYRl
Dca27529.1 211 KPRVVWSVELHQQFVAAVNQL-GIDKAVPKKILELMNVPGLTRENVASHLQKYRL 264
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 73.8 | 6.7e-25 | 36 | 144 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTTES CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealkaGak 95
vl+vdD+p+ + +l+++l+ +y +v+ + +e al ll+e++ +D+++ D+ mp+mdG++ll++ e +lp+i+++a ++++ + + + Ga
Dca27529.1 36 VLVVDDDPTCLIILEKMLRTCRY-DVTKTNRAEFALSLLRENKngFDIVISDVHMPDMDGFKLLEHVGLEM-DLPVIMMSADDSKSVVMKGVTHGAC 130
89*********************.***************999999**********************6644.8************************ PP
EEEESS--HHHHHH CS
Response_reg 96 dflsKpfdpeelvk 109
d+l Kp+ +e+l +
Dca27529.1 131 DYLIKPVRIEALKN 144
**********9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 6.2E-170 | 11 | 639 | IPR017053 | Response regulator B-type, plant |
| SuperFamily | SSF52172 | 5.55E-34 | 33 | 156 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 9.0E-41 | 33 | 169 | No hit | No description |
| SMART | SM00448 | 1.8E-27 | 34 | 146 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 40.954 | 35 | 150 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 6.6E-22 | 36 | 144 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 9.42E-25 | 37 | 149 | No hit | No description |
| PROSITE profile | PS51294 | 11.048 | 208 | 267 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.4E-20 | 208 | 268 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-30 | 210 | 269 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.1E-25 | 211 | 264 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.2E-8 | 213 | 263 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 648 aa Download sequence Send to blast |
MNLGGGLMGS MSLPSSTMTR KPVDGGSDPF PAGLRVLVVD DDPTCLIILE KMLRTCRYDV 60 TKTNRAEFAL SLLRENKNGF DIVISDVHMP DMDGFKLLEH VGLEMDLPVI MMSADDSKSV 120 VMKGVTHGAC DYLIKPVRIE ALKNIWQHVV RKKKNEWRDV EQSSSYDEGD RQQKQDDVVS 180 SPANEGSWKS SKRKAEDDDA DEKDDTSTLK KPRVVWSVEL HQQFVAAVNQ LGIDKAVPKK 240 ILELMNVPGL TRENVASHLQ KYRLYLRRLS GVSQHQGSGL NGSFMSQDGG FGAMSQLSGI 300 DLQALHATAQ LSSQTLAAYT RLPPIKPSMS MPFADQRNLF SFENTATSRY VDGQQQQQQY 360 SKQLNLLHGI PTTLESKQLA GLNQSAQVAG LNQSPQIAGL NQSAQISKNV NLQGSASSSH 420 HTSSLLMPVV PQHSRGQVLN ENVTGHVSRT QPSNRGYNRV SQAGRLVDYS INQIPDLPGT 480 TSFPLGSTPG LTSLASKGVF NQEDDSHDIK AMRGYVGSYD VFPEAHKAQD WQVRNPNMVY 540 GVSGPSSKLP TSVQSSINVT PGNPMFSSGL DNHHLGMTNI SHNGNGILVD STPRVKAEKI 600 SDVVSPGTNL FTEHYTQDDL MSALFKQHDG IGSGEYDFDG YSLDNIPV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 3e-22 | 210 | 270 | 4 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-[AG]GATT-3'. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. Could directly activate some type-A response regulators in response to cytokinins. Involved in the expression of nuclear genes for components of mitochondrial complex I. Promotes cytokinin-mediated leaf longevity. Involved in the ethylene signaling pathway in an ETR1-dependent manner and in the cytokinin signaling pathway. {ECO:0000269|PubMed:11370868, ECO:0000269|PubMed:11574878, ECO:0000269|PubMed:15282545, ECO:0000269|PubMed:16407152}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
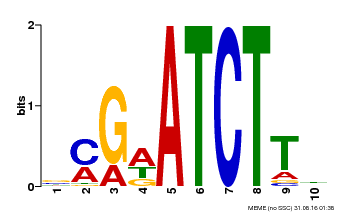
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010670915.1 | 0.0 | PREDICTED: two-component response regulator ARR2 isoform X2 | ||||
| Swissprot | Q9ZWJ9 | 1e-177 | ARR2_ARATH; Two-component response regulator ARR2 | ||||
| TrEMBL | A0A0K9RPY9 | 0.0 | A0A0K9RPY9_SPIOL; Two-component response regulator | ||||
| STRING | XP_010670914.1 | 0.0 | (Beta vulgaris) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G16857.2 | 1e-171 | response regulator 1 | ||||



