 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | DCAR_023184 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; campanulids; Apiales; Apiineae; Apiaceae; Apioideae; Scandiceae; Daucinae; Daucus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 303aa MW: 32840 Da PI: 7.2927 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 26.8 | 1.2e-08 | 11 | 55 | 4 | 45 |
S-HHHHHHHHHHHHHTTTT....-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 4 WTteEdellvdavkqlGgg....tWktIartmgkgRtlkqcksrwq 45
W+ eE++ + +a++++ + +W++Ia +++ +++ +q+ +++
DCAR_023184 11 WSREEEKCFENAIAMHWIEdskeQWDKIATMVP-TKSVEQLHEHYK 55
****************99999************.**********97 PP
| |||||||
| 2 | Myb_DNA-binding | 37 | 7.7e-12 | 143 | 187 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ l++ + ++G+g+W++I+r Rt+ q+ s+ qky
DCAR_023184 143 PWTEEEHRLFLIGLDKFGKGDWRSISRNYVISRTPTQVASHAQKY 187
7*****************************89************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 11.703 | 6 | 62 | IPR017884 | SANT domain |
| SMART | SM00717 | 3.5E-5 | 7 | 60 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.3E-4 | 10 | 57 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 2.23E-9 | 11 | 65 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 9.0E-7 | 11 | 55 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.24E-8 | 11 | 58 | No hit | No description |
| PROSITE profile | PS51294 | 17.889 | 136 | 192 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.48E-17 | 138 | 193 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 8.0E-9 | 140 | 190 | IPR001005 | SANT/Myb domain |
| TIGRFAMs | TIGR01557 | 8.9E-17 | 140 | 190 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 3.4E-10 | 141 | 186 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 8.11E-9 | 143 | 188 | No hit | No description |
| Pfam | PF00249 | 2.9E-10 | 143 | 187 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 303 aa Download sequence Send to blast |
MSDSGSGSEM WSREEEKCFE NAIAMHWIED SKEQWDKIAT MVPTKSVEQL HEHYKLLVED 60 VNAIEAGLVP LPKYAGEEEA AAAAAAVSSS STKSNHQQGL MNATAKRSSC NFSSGFSAFG 120 GDSTCLGSKG SSRLEQERRK GIPWTEEEHR LFLIGLDKFG KGDWRSISRN YVISRTPTQV 180 ASHAQKYFIR LNSMNRDRRR TSIHDITSVN NGDVSSQHIP ITGQQLTSDP SNGMAVGPPM 240 KHPGPPGMAM YGAPLGHPVV AGSAHLGSAV GTPVMLPPPI HHHHPYVLPV AYPMAPPPAM 300 HQ* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that coordinates abscisic acid (ABA) biosynthesis and signaling-related genes via binding to the specific promoter motif 5'-(A/T)AACCAT-3'. Represses ABA-mediated salt (e.g. NaCl and KCl) stress tolerance. Regulates leaf shape and promotes vegetative growth. {ECO:0000269|PubMed:26243618}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00191 | DAP | Transfer from AT1G49010 | Download |
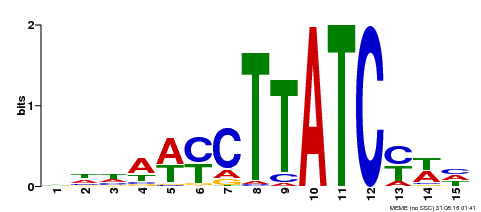
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salicylic acid (SA) and gibberellic acid (GA) (PubMed:16463103). Triggered by dehydration and salt stress (PubMed:26243618). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:26243618}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017257768.1 | 0.0 | PREDICTED: transcription factor DIVARICATA | ||||
| Swissprot | Q9FNN6 | 6e-81 | SRM1_ARATH; Transcription factor SRM1 | ||||
| TrEMBL | A0A161XA29 | 0.0 | A0A161XA29_DAUCS; Uncharacterized protein | ||||
| STRING | VIT_17s0000g04130.t01 | 1e-132 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA5801 | 21 | 36 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G49010.1 | 2e-82 | MYB family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | DCAR_023184 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



