 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | DCAR_017224 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; campanulids; Apiales; Apiineae; Apiaceae; Apioideae; Scandiceae; Daucinae; Daucus
|
||||||||
| Family | BES1 | ||||||||
| Protein Properties | Length: 315aa MW: 33943.1 Da PI: 9.1157 | ||||||||
| Description | BES1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF822 | 211.2 | 2.7e-65 | 5 | 149 | 1 | 149 |
DUF822 1 ggsgrkptwkErEnnkrRERrRRaiaakiyaGLRaqGnyklpkraDnneVlkALcreAGwvvedDGttyrkgskpleeaeaagssasaspesslqssl 98
g++gr ptwkErEnnkrRERrRRai+akiyaGLRaqGn+klpk++DnneVlkALc+eAGw+v++DGttyrkg+kp+ + +g+ +++s +ss+q s+
DCAR_017224 5 GATGRLPTWKERENNKRRERRRRAISAKIYAGLRAQGNFKLPKHCDNNEVLKALCNEAGWIVQEDGTTYRKGCKPPPGEILTGTPTNMSACSSMQPSP 102
5899************************************************************************8888899999************ PP
DUF822 99 kssalaspvesysaspksssfpspssldsislasaasllpvlsvlslvsss 149
ssa++sp++sy+asp+sssfpsp++ + s + +lp+l +l++++ss
DCAR_017224 103 MSSAFPSPAPSYHASPTSSSFPSPTRETNPS----SYILPFLCNLAAIPSS 149
*************************966553....4688888888877776 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05687 | 1.7E-62 | 7 | 130 | IPR008540 | BES1/BZR1 plant transcription factor, N-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009742 | Biological Process | brassinosteroid mediated signaling pathway | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048316 | Biological Process | seed development | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 315 aa Download sequence Send to blast |
MTAGGATGRL PTWKERENNK RRERRRRAIS AKIYAGLRAQ GNFKLPKHCD NNEVLKALCN 60 EAGWIVQEDG TTYRKGCKPP PGEILTGTPT NMSACSSMQP SPMSSAFPSP APSYHASPTS 120 SSFPSPTRET NPSSYILPFL CNLAAIPSSL PPLRISNSAP VTPPLSSPTA RGSKRPEWEL 180 PTNNSMLFRH PLFAASAPAS PTRRHHHHHF TPATIPECDE SDASTVDSGR WVSFQPSPSQ 240 AVPTSPTFNL VKPVAHQIGA VDTHGGLGWG AAAKRARGPE FEFESCTVRA WEGERIHEVG 300 VDDLELTLGS GKAH* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zd4_A | 8e-28 | 9 | 84 | 372 | 447 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_B | 8e-28 | 9 | 84 | 372 | 447 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_C | 8e-28 | 9 | 84 | 372 | 447 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_D | 8e-28 | 9 | 84 | 372 | 447 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| Search in ModeBase | ||||||
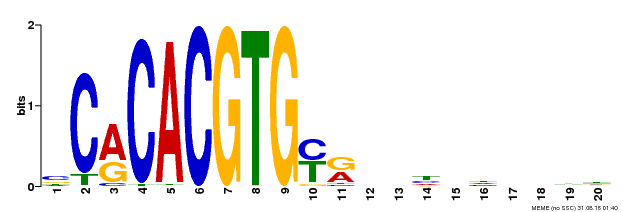
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00237 | DAP | Transfer from AT1G75080 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_017253773.1 | 0.0 | PREDICTED: BES1/BZR1 homolog protein 2 | ||||
| Swissprot | Q94A43 | 1e-109 | BEH2_ARATH; BES1/BZR1 homolog protein 2 | ||||
| TrEMBL | A0A161YKM1 | 0.0 | A0A161YKM1_DAUCS; Uncharacterized protein | ||||
| STRING | XP_009349751.1 | 1e-142 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA9486 | 21 | 28 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G75080.2 | 4e-89 | BES1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | DCAR_017224 |



