 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | orange1.1g018802m | ||||||||
| Common Name | CISIN_1g018802mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Sapindales; Rutaceae; Aurantioideae; Citrus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 351aa MW: 38682 Da PI: 8.0724 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 100.9 | 8.1e-32 | 145 | 198 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
pr+rWt++LH++Fv+av+ LGG+e+AtPk++lelm+vk+Ltl+hvkSHLQ+YR+
orange1.1g018802m 145 PRMRWTTTLHAHFVHAVQLLGGHERATPKSVLELMNVKDLTLAHVKSHLQMYRT 198
9****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.97E-15 | 142 | 199 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 4.5E-28 | 143 | 199 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.3E-23 | 145 | 198 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.3E-7 | 146 | 197 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005618 | Cellular Component | cell wall | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 351 aa Download sequence Send to blast |
MFSSMNPLIR TASLSPMPDL SLQISPPSAS DCEANNNKEV RFDGLVRNSY AFYSDGGSTT 60 DSASSGSDLS HENRFYNPAE RSYNLGPSEP MLSLGFEMAD LCPPPPQQLP RTINHHQQYQ 120 PQIHGHEFKR NARMISGVKR SVRAPRMRWT TTLHAHFVHA VQLLGGHERA TPKSVLELMN 180 VKDLTLAHVK SHLQMYRTVK STDKGSGQGQ TDMGLNQRTG VVDLEGGLSC AKSDTNSSLH 240 PTPSSPQATS QQKRQRGSLP SMETNNRSIS NSGNAMTYSH FKANDTKGDG RKTAVHMSDN 300 NKVEVERLDS SSLTASDDKL LNLEFTLGRP SWQMDYADQS SNHELNLLHC * |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 5e-17 | 146 | 200 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 5e-17 | 146 | 200 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 5e-17 | 146 | 200 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 5e-17 | 146 | 200 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00107 | PBM | Transfer from AT5G42630 | Download |
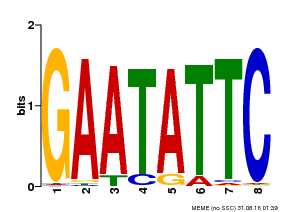
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KP271989 | 6e-94 | KP271989.1 UNVERIFIED: Populus trichocarpa isolate Potri.014G037200_BESC-125 mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006434889.1 | 0.0 | probable transcription factor KAN4 | ||||
| TrEMBL | A0A067H0C9 | 0.0 | A0A067H0C9_CITSI; Uncharacterized protein | ||||
| STRING | XP_006434889.1 | 0.0 | (Citrus clementina) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM6572 | 28 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G42630.1 | 1e-69 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | orange1.1g018802m |



