 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.212820.1 | ||||||||
| Common Name | Csa_1G467710, LOC101204390 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 410aa MW: 46704.4 Da PI: 4.9535 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 108.7 | 4.4e-34 | 15 | 105 | 2 | 101 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkellek 96
Fl k+y++++d+++++++sws++ +sfvv+++ ef++ +Lpk+Fkhsnf+SF+RQLn+YgF+kv+ e+ weF+++ F +gk +l+++
Cucsa.212820.1 15 FLVKTYDMVDDPSTNSIVSWSSSDKSFVVWNPLEFSSVLLPKFFKHSNFSSFIRQLNTYGFRKVDPEQ---------WEFSNEDFVRGKPHLMKN 100
9********************999****************************************9998.........****************** PP
XXXXX CS
HSF_DNA-bind 97 ikrkk 101
i+r+k
Cucsa.212820.1 101 IHRRK 105
***98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00415 | 2.0E-59 | 11 | 104 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| SuperFamily | SSF46785 | 7.62E-34 | 11 | 104 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Gene3D | G3DSA:1.10.10.10 | 2.9E-37 | 12 | 104 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| PRINTS | PR00056 | 6.2E-19 | 15 | 38 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Pfam | PF00447 | 7.2E-31 | 15 | 104 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 6.2E-19 | 53 | 65 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 54 | 78 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 6.2E-19 | 66 | 78 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 410 aa Download sequence Send to blast |
MDEAQGGGLT SLPPFLVKTY DMVDDPSTNS IVSWSSSDKS FVVWNPLEFS SVLLPKFFKH 60 SNFSSFIRQL NTYGFRKVDP EQWEFSNEDF VRGKPHLMKN IHRRKPIHSH SLQNLHGQGI 120 SPLTEVERNS FKDDIERLKL DKEQLLLELQ KYEQEYQGVG LQMQNLKDQF QRVQQEMQLF 180 ISLMARLLQK PGLHLDLLPQ LETPERKRRL PRVSYNISED SLEDNHLGTT QTIGRDDMGC 240 SFDPILEKEQ LELLETSLTF WEGIIHSYDE TVSPLDSSSN LELVGSVSHA SSPAISCRLV 300 REEFRCKSPG IDMNLEPMAT VAPDSVASKE QAAGVNAPLP TGFNDVFWQQ FLTENPGASD 360 PQEVQSARKD SDVINEENRQ SDHGKFWWNT RSVNNVVEQI GHLKPAEKF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 1e-21 | 8 | 104 | 22 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 1e-21 | 8 | 104 | 22 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 1e-21 | 8 | 104 | 22 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 204 | 209 | ERKRRL |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00443 | DAP | Transfer from AT4G18880 | Download |
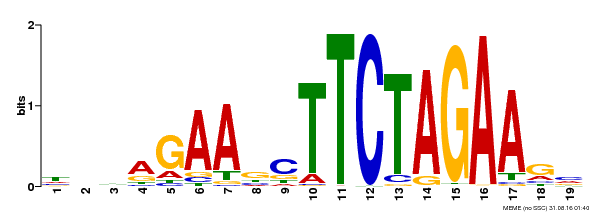
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681932 | 0.0 | LN681932.1 Cucumis melo genomic scaffold, anchoredscaffold00001. | |||
| GenBank | LN713266 | 0.0 | LN713266.1 Cucumis melo genomic chromosome, chr_12. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011657567.1 | 0.0 | PREDICTED: heat stress transcription factor A-4c-like | ||||
| Refseq | XP_011657570.1 | 0.0 | PREDICTED: heat stress transcription factor A-4c-like | ||||
| TrEMBL | A0A0A0LV22 | 0.0 | A0A0A0LV22_CUCSA; Uncharacterized protein | ||||
| STRING | XP_004161196.1 | 0.0 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3568 | 33 | 61 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G18880.1 | 1e-103 | heat shock transcription factor A4A | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.212820.1 |
| Entrez Gene | 101204390 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



