 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.139290.1 | ||||||||
| Common Name | Csa_6G185260 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | C3H | ||||||||
| Protein Properties | Length: 668aa MW: 72566.9 Da PI: 6.6841 | ||||||||
| Description | C3H family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-CCCH | 21.3 | 4.6e-07 | 259 | 279 | 5 | 26 |
--SGGGGTS--TTTTT-SS-SS CS
zf-CCCH 5 lCrffartGtCkyGdrCkFaHg 26
+C+ f++ GtC+ Gd C +aHg
Cucsa.139290.1 259 PCPEFRK-GTCRQGDACEYAHG 279
8******.*************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF48403 | 3.11E-13 | 56 | 134 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 16.175 | 59 | 134 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.0049 | 59 | 89 | IPR002110 | Ankyrin repeat |
| Pfam | PF12796 | 8.1E-7 | 59 | 126 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.2E-14 | 60 | 139 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.36E-12 | 61 | 158 | No hit | No description |
| PROSITE profile | PS50088 | 11.648 | 94 | 129 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.23 | 94 | 126 | IPR002110 | Ankyrin repeat |
| SMART | SM00356 | 5.4E-5 | 254 | 280 | IPR000571 | Zinc finger, CCCH-type |
| Pfam | PF00642 | 1.2E-4 | 259 | 279 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 12.784 | 259 | 281 | IPR000571 | Zinc finger, CCCH-type |
| Gene3D | G3DSA:4.10.1000.10 | 8.5E-4 | 287 | 317 | IPR000571 | Zinc finger, CCCH-type |
| SMART | SM00356 | 30 | 289 | 312 | IPR000571 | Zinc finger, CCCH-type |
| PROSITE profile | PS50103 | 7.103 | 289 | 313 | IPR000571 | Zinc finger, CCCH-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 668 aa Download sequence Send to blast |
MDGGFKRGGL GFCISVLLEF SATDDLIGFR SAVEEDGHDI DEASLWYGRI FGSKKMGYEE 60 RTPLMVAAMF GSLNVLSYIL HSGRVDVNRA CGSDGVTTLH CAVAGGSAVV DQVVKLLLDA 120 SADVSAVDAN GNRPGDLIAP DFTSAFYSRK KTLQQLLNGH EGLSSSEAIF YERETLEPLE 180 LSTLRASRDG TEKKEYPVDL SLPDIKNGIY STDEFRMYTF KIKPCTRAYS HDWTECPFVH 240 PGENARRRDP RKYHYSCVPC PEFRKGTCRQ GDACEYAHGI FECWLHPAQY RTRLCKDETG 300 CTRKVCFFAH KPEELRPLYA STGSAVLSPR SICGSSLDIA SISSLTLGSP SALIPPSSTP 360 PLTPSGVSSP MGGTMWQTQC NIAPPTLHLP GSRLKASLSA RDVDLDVELL GLESQRRRQQ 420 QLMDEMSCLS SPSRWNNGLP TPASFPSPRS RNGELNGLGG MKQTNLEDFF GSVDPAILPQ 480 LQGLSLDSVG SQVQSPSGIQ MRQSLNQSFL SSYGNSIGSP PPRLSQPSVS TAASVLSSRA 540 AAFAKRSQSF IERSMVSRHT GLSPPGTSTT AMPLNLSDWG SPDGKLDWGI RGEELNKLKK 600 SASFGIRNNC TSSPVTSTMH TTAPEPDVSW VQSLVKDAPS ENAVQLKYCI QSTINQIWAE 660 EMFLPIP* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00566 | DAP | Transfer from AT5G58620 | Download |
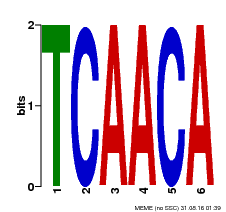
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681919 | 0.0 | LN681919.1 Cucumis melo genomic scaffold, anchoredscaffold00084. | |||
| GenBank | LN713265 | 0.0 | LN713265.1 Cucumis melo genomic chromosome, chr_11. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004143161.1 | 0.0 | PREDICTED: zinc finger CCCH domain-containing protein 66 | ||||
| Swissprot | Q9LUZ4 | 0.0 | C3H66_ARATH; Zinc finger CCCH domain-containing protein 66 | ||||
| TrEMBL | A0A0A0KH40 | 0.0 | A0A0A0KH40_CUCSA; Uncharacterized protein | ||||
| STRING | XP_004143161.1 | 0.0 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1796 | 34 | 90 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G58620.1 | 0.0 | C3H family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.139290.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



