 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cucsa.112760.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 334aa MW: 37791.5 Da PI: 8.5402 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 55.3 | 1.5e-17 | 15 | 62 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd +lv +++++G+g+W++++ g+ R+ k+c++rw +yl
Cucsa.112760.1 15 KGPWTPEEDIILVSYIQEHGPGNWRSVPTNTGLLRCSKSCRLRWTNYL 62
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 48.4 | 2.2e-15 | 68 | 113 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T E++ ++++ ++lG++ W++Ia++++ Rt++++k++w+++l
Cucsa.112760.1 68 RGNFTDHEEKMIIHLQALLGNR-WAAIASYLP-QRTDNDIKNYWNTHL 113
89********************.*********.************996 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-23 | 6 | 65 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 24.836 | 10 | 66 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.86E-30 | 12 | 109 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.4E-14 | 14 | 64 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.3E-16 | 15 | 62 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.60E-10 | 17 | 62 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.7E-24 | 66 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.0E-15 | 67 | 115 | IPR001005 | SANT/Myb domain |
| PROSITE profile | PS51294 | 19.216 | 67 | 117 | IPR017930 | Myb domain |
| Pfam | PF00249 | 8.6E-14 | 68 | 113 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.97E-10 | 70 | 113 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0080167 | Biological Process | response to karrikin | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 334 aa Download sequence Send to blast |
MMGRPPCCDK IGVKKGPWTP EEDIILVSYI QEHGPGNWRS VPTNTGLLRC SKSCRLRWTN 60 YLRPGIKRGN FTDHEEKMII HLQALLGNRW AAIASYLPQR TDNDIKNYWN THLKKKLRKM 120 QSLGGGGGSE GSGVQSQSNT TLSKGQWERR LQTDIHTAKK ALCEALSLEK KPDLEEYYYN 180 NNLKILQESP TAPTPPPQTI TTTYASSAEN IARLLENWMK KSPTKSSEIT TTTQVCMSEE 240 GSQISATNNY NNNSPDHQNP NNDNNNNLEG FFNNYSSHST TNNQEVVPLT FLEKWLFDDC 300 HGMRVFGLLE SFCFSFVYMM TPPQTQQKNK SVN* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 3e-25 | 15 | 117 | 27 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. {ECO:0000305}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00395 | DAP | Transfer from AT3G47600 | Download |
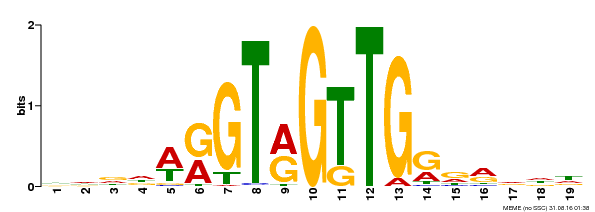
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681932 | 0.0 | LN681932.1 Cucumis melo genomic scaffold, anchoredscaffold00001. | |||
| GenBank | LN713266 | 0.0 | LN713266.1 Cucumis melo genomic chromosome, chr_12. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004137409.1 | 0.0 | PREDICTED: myb-related protein 306 | ||||
| Swissprot | P81392 | 1e-116 | MYB06_ANTMA; Myb-related protein 306 | ||||
| TrEMBL | A0A1S3B5N8 | 0.0 | A0A1S3B5N8_CUCME; myb-related protein 306-like | ||||
| STRING | XP_004137409.1 | 0.0 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1016 | 34 | 115 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G47600.1 | 1e-105 | myb domain protein 94 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cucsa.112760.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



