 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK28233.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1057aa MW: 119109 Da PI: 6.0341 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 178.2 | 1e-55 | 20 | 136 | 2 | 118 |
CG-1 2 lkekkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywl 101
++ ++rwl++ ei++iL n++k++++ e+++ p++gsl+L++rk++ryfrkDG++w+kkkdgktv+E+he+LK g+++vl+cyYah+een++fqrr+yw+
PK28233.1 20 IEAQHRWLRPAEICEILRNYKKFRIAPEPANMPPNGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHERLKAGSIDVLHCYYAHGEENENFQRRSYWM 119
5679************************************************************************************************ PP
CG-1 102 Leeelekivlvhylevk 118
Lee+l++ivlvhy++vk
PK28233.1 120 LEEDLSHIVLVHYRQVK 136
**************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 81.586 | 15 | 141 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 4.7E-77 | 18 | 136 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 5.6E-49 | 21 | 134 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 3.1E-5 | 471 | 558 | IPR013783 | Immunoglobulin-like fold |
| Pfam | PF01833 | 3.6E-6 | 473 | 557 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 4.9E-17 | 473 | 558 | IPR014756 | Immunoglobulin E-set |
| PROSITE profile | PS50297 | 17.024 | 638 | 777 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 8.2E-18 | 651 | 800 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 9.79E-12 | 654 | 765 | No hit | No description |
| SuperFamily | SSF48403 | 2.64E-16 | 662 | 768 | IPR020683 | Ankyrin repeat-containing domain |
| Pfam | PF12796 | 2.2E-6 | 678 | 769 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.041 | 706 | 735 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.084 | 706 | 738 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 840 | 745 | 774 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.088 | 879 | 901 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 1.09E-7 | 880 | 930 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| PROSITE profile | PS50096 | 8.682 | 881 | 909 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0035 | 882 | 900 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0026 | 902 | 924 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.761 | 903 | 927 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 1.7E-4 | 905 | 924 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0050832 | Biological Process | defense response to fungus | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005516 | Molecular Function | calmodulin binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1057 aa Download sequence Send to blast |
MADTRRLGLP NQLDIEQILI EAQHRWLRPA EICEILRNYK KFRIAPEPAN MPPNGSLFLF 60 DRKVLRYFRK DGHNWRKKKD GKTVKEAHER LKAGSIDVLH CYYAHGEENE NFQRRSYWML 120 EEDLSHIVLV HYRQVKGNRT HYNRIKGTEE ADTAPSSEID SPASSSIHPN SYQMISQTTD 180 TTSLNSAQAS EYEDAESAYN QASSRLHSFL DLQQPFAEKI SAGINDAYYP VSFSNNNQEK 240 LSDIPGMDSS AGVSSGHSKN LDFPAWDNTL GNNAASIHLP FQQAFSSSQS NNVTVIQKQE 300 QGPFEQLFSN GFGKHPQIQE DWQASEGNSR LPEWSVDQTL LTGSDNNLTS RYRGEVAHGD 360 ELLKIHRGNI ERDDFVKPVS KSNMNLEEKP YNSGTKQPLL VGSFAEEGLK KLDSFNRWMS 420 KELGDVNESH MQTSSRAYWD SVEGDTCVND SSQVRLDNYV LSPSLSQEQL FSIIDFSPNW 480 AYESSEVKVL ITGRFLKVEQ AEINKWSCMF GEVEVPTEII AEGVLRCNAP IHKAGRVPFY 540 VTCSNRLACS EVREFEYRLS EARDVQTKYN QSGCTNEILK LRFGKLMSLS SPSSNADPVN 600 LAEMSQLSNK ISSLLKEDQD EWDQMYRLTS EENFSVESVQ EQLHQKLLKE KLYEWLLQKV 660 AEGGKGPSLL DEGGQGVLHF AAALGYDWAL EPTIVAGVSV NFRDVNGWTA LHWAAFCGRE 720 RTVASLISLG AAPGLLTDPS PKYPSGKTPA DLASDNGHKG ISGYLAESAL SAHLKCLDLD 780 AKASKATDTS TPNAVQTVSE RTATPIKDDD PDRLSLKDSL AAVCNATQAA ARIHQVFRVQ 840 SFQRKQLEEY GDDKFGLSDE RALSLIAVKT QKGGQHNNDV NAAAIRIQNK FRGWKGRKEF 900 LSIRQNIVKI QALVRGHQVR KNYKKFVWSV GIVEKIILRW RRKGSGLRGF KSEALVEVPS 960 IEDPSSSKDD DYDFLKQGRK QSEERMQKAL ARVKSMVQYP EARNQYRRLL NVVTEIQEKK 1020 VPYDDVMNTE GTIDFEDDLI DIEALLEDDT YMPTAIQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cxk_A | 2e-13 | 473 | 558 | 9 | 89 | calmodulin binding transcription activator 1 |
| 2cxk_B | 2e-13 | 473 | 558 | 9 | 89 | calmodulin binding transcription activator 1 |
| 2cxk_C | 2e-13 | 473 | 558 | 9 | 89 | calmodulin binding transcription activator 1 |
| 2cxk_D | 2e-13 | 473 | 558 | 9 | 89 | calmodulin binding transcription activator 1 |
| 2cxk_E | 2e-13 | 473 | 558 | 9 | 89 | calmodulin binding transcription activator 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3'. Binds calmodulin in a calcium-dependent manner in vitro (PubMed:12218065). Regulates transcriptional activity in response to calcium signals (Probable). Involved in freezing tolerance in association with CAMTA1 and CAMTA2 (PubMed:23581962). Required for the cold-induced expression of DREB1B/CBF1, DREB1C/CBF2, ZAT12 and GOLS3 (PubMed:19270186). Involved in response to cold. Contributes together with CAMTA5 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:28351986). Involved together with CAMTA2 and CAMTA4 in the positive regulation of a general stress response (GSR) (PubMed:25039701). Involved in the regulation of GSR amplitude downstream of MEKK1 (PubMed:25157030). Involved in the regulation of a set of genes involved in defense responses against pathogens (PubMed:18298954). Involved in the regulation of both basal resistance and systemic acquired resistance (SAR) (PubMed:21900483). Acts as negative regulator of plant immunity (PubMed:19122675, PubMed:21900483, PubMed:22345509, PubMed:28407487). Binds to the promoter of the defense-related gene EDS1 and represses its expression (PubMed:19122675). Binds to the promoter of the defense-related gene NDR1 and represses its expression (PubMed:22345509). Involved in defense against insects (PubMed:23072934, PubMed:22371088). Required for tolerance to the generalist herbivore Trichoplusia ni, and contributes to the positive regulation of genes associated with glucosinolate metabolism (PubMed:23072934). Required for tolerance to Bradysia impatiens larvae. Mediates herbivore-induced wound response (PubMed:22371088). Required for wound-induced jasmonate accumulation (PubMed:23072934, PubMed:22371088). Involved in the regulation of ethylene-induced senescence by binding to the promoter of the senescence-inducer gene EIN3 and repressing its expression (PubMed:22345509). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:18298954, ECO:0000269|PubMed:19122675, ECO:0000269|PubMed:19270186, ECO:0000269|PubMed:21900483, ECO:0000269|PubMed:22345509, ECO:0000269|PubMed:22371088, ECO:0000269|PubMed:23072934, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25157030, ECO:0000269|PubMed:28351986, ECO:0000269|PubMed:28407487, ECO:0000305|PubMed:11925432}. | |||||
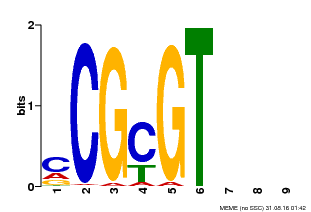
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00042 | PBM | Transfer from AT2G22300 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By heat shock, UVB, salt, wounding, ethylene and methyl jasmonate (PubMed:11162426, PubMed:12218065). Induced by infection with the fungal pathogen Golovinomyces cichoracearum (powdery mildew) and the bacterial pathogen Pseudomonas syringae pv tomato strain DC3000 (PubMed:22345509). {ECO:0000269|PubMed:11162426, ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:22345509}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024021686.1 | 0.0 | calmodulin-binding transcription activator 3 | ||||
| Swissprot | Q8GSA7 | 0.0 | CMTA3_ARATH; Calmodulin-binding transcription activator 3 | ||||
| TrEMBL | A0A2P5BFW1 | 0.0 | A0A2P5BFW1_PARAD; Notch | ||||
| STRING | XP_010087599.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8284 | 27 | 35 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G22300.2 | 0.0 | signal responsive 1 | ||||



