 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK27491.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 489aa MW: 52581.6 Da PI: 6.0887 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 87.7 | 1.1e-27 | 198 | 252 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k +++WtpeLH+rFv+aveqL G +kA+P++ilelm++++Lt+++++SHLQkYR++
PK27491.1 198 KVKVDWTPELHRRFVQAVEQL-GVDKAVPSRILELMGIDCLTRHNIASHLQKYRSH 252
5689*****************.********************************86 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 18.152 | 195 | 254 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 3.94E-18 | 196 | 255 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.4E-26 | 196 | 255 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.2E-26 | 198 | 252 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 5.5E-8 | 201 | 250 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007165 | Biological Process | signal transduction | ||||
| GO:0009658 | Biological Process | chloroplast organization | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0010380 | Biological Process | regulation of chlorophyll biosynthetic process | ||||
| GO:0010638 | Biological Process | positive regulation of organelle organization | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 489 aa Download sequence Send to blast |
MIIGMLAVSS LRDTYKDENQ GGEAMETSNH INFSINGGDG FPDFADCIDF DDLFIGINVD 60 GDVLPDLEMD PEILADFSAE DYNSDDMNTS PLSAEKSADI DDNNVSSINS AITSTTTTTT 120 TTTHAEEEDN NNNRKIKSTT AETEDDKDSG SGSGSVTGGD VASANNNKEA SDNKGGGKKS 180 SSSTTAQSKN NNSQGKRKVK VDWTPELHRR FVQAVEQLGV DKAVPSRILE LMGIDCLTRH 240 NIASHLQKYR SHRKHLLARE AEAASWSQRR QMFGAAPGGG GGAVAGKRDV SPWAVAPTMG 300 FPQQAVVPQP PPVPMHPHFR PLHVWGHPTV DQSLMHVWPP KHLPHSPSHH PPPPPPAWQP 360 AVPTPPPLHD PSYWHSHPHR VPSALTPGTP CFPQPMATPR FSTPAVPGIP PGHAMYKVDA 420 GIGNLTPGQS GPHPLFDFHP SKESVDEAIG DVLAKPWLPL PLGLKPPSMD SVMGELQRQG 480 VPRIPPSCA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 3e-16 | 195 | 250 | 2 | 57 | ARR10-B |
| Search in ModeBase | ||||||
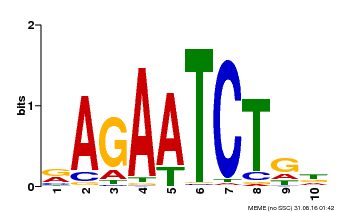
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00022 | PBM | Transfer from AT2G20570 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010093955.1 | 1e-162 | transcription activator GLK1 isoform X2 | ||||
| TrEMBL | A0A2P5B6N0 | 0.0 | A0A2P5B6N0_PARAD; Octamer-binding transcription factor | ||||
| STRING | XP_010093955.1 | 1e-162 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4242 | 34 | 61 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G20570.1 | 9e-79 | GBF's pro-rich region-interacting factor 1 | ||||



