 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK19114.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | EIL | ||||||||
| Protein Properties | Length: 592aa MW: 66761.6 Da PI: 5.4513 | ||||||||
| Description | EIL family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | EIN3 | 434.1 | 1.3e-132 | 33 | 361 | 1 | 315 |
XXXXXXXXXXXXXXXXXXXXXXX..XXXXX.XXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXXX CS
EIN3 1 eelkkrmwkdqmllkrlkerkkqlledkeaatgakksnksneqarrkkmsraQDgiLkYMlkemevcnaqGfvYgiipekgkpvegasdsLraWWkekve 100
eel++rmwkd+++lkrlker+k +++ a++++k +++++qarrkkmsraQDgiLkYMlk mevc+a+GfvYgiipekgkpv+gasd++raWWkekv+
PK19114.1 33 EELERRMWKDRIKLKRLKERQKLA--AQQ-AAEKQKPKQTSDQARRKKMSRAQDGILKYMLKLMEVCKARGFVYGIIPEKGKPVSGASDNIRAWWKEKVK 129
8*******************9963..344.599******************************************************************* PP
XXXXXXXXXXXXXXXXXXXXXXXXX....XX----STTS-HHHHHHHHHHHSSSSSS-TTS--TTT--HHHH---S--HHHHHHT--TT--.-----GGG CS
EIN3 101 fdrngpaaiskyqaknlilsgesslqtersseshslselqDTtlgSLLsalmqhcdppqrrfplekgvepPWWPtGkelwwgelglskdqgtppykkphd 200
fd+ngpaai+ky+a++l++ +++++ ++ ++++ l++lqD+tlgSLLs+lmqhc+ppqr+fplek v+pPWWPtGke+ww +lgl++ q ppykkphd
PK19114.1 130 FDKNGPAAIAKYEAECLAMCEADNN--RNGNSQSILQDLQDATLGSLLSSLMQHCEPPQRKFPLEKKVPPPWWPTGKEDWWVKLGLPHVQT-PPYKKPHD 226
******************9988888..68999999****************************************************9887.******** PP
--HHHHHHHHHHHHHHTGGGHHHHHHTTTTSSSSTTT--SHHHHHHHHHHTTTTT-S--XXXXXX.......XXXXXXXXXXXXXXXXXXXXXXXXXX.X CS
EIN3 201 lkkawkvsvLtavikhmsptieeirelerqskylqdkmsakesfallsvlnqeekecatvsahss.......slrkqspkvtlsceqkedvegkkeskik 293
lkk+wkv+vLtavikhmsp+i++ir+++rqsk+lqdkm+akes+++l vl++ee+++++ s+++ +++++vt+++++++dv+g ++ +
PK19114.1 227 LKKMWKVGVLTAVIKHMSPDIAKIRRHVRQSKCLQDKMTAKESAIWLGVLSREEALIRQPSSDNGmsgvteiPPGRGEKQVTANSDSNYDVDGADDGLGS 326
***************************************************************44446766645569***************77776656 PP
XXXXXXXXX.......................XXXXXXXXXXXXX CS
EIN3 294 hvqavktta.......................gfpvvrkrkkkps 315
++ +++++ + p+
PK19114.1 327 VSSRDERRSqsmdvepsgsqqnntsqpgqdkeK----------PD 361
555554444555555566665533333333332..........33 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF04873 | 8.7E-126 | 33 | 280 | No hit | No description |
| Gene3D | G3DSA:1.10.3180.10 | 1.1E-70 | 154 | 288 | IPR023278 | Ethylene insensitive 3-like protein, DNA-binding domain |
| SuperFamily | SSF116768 | 4.58E-61 | 161 | 284 | IPR023278 | Ethylene insensitive 3-like protein, DNA-binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0042762 | Biological Process | regulation of sulfur metabolic process | ||||
| GO:0071281 | Biological Process | cellular response to iron ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 592 aa Download sequence Send to blast |
MAYDGDSSDI EVDDLRCDDI ADKDVSDEEI EPEELERRMW KDRIKLKRLK ERQKLAAQQA 60 AEKQKPKQTS DQARRKKMSR AQDGILKYML KLMEVCKARG FVYGIIPEKG KPVSGASDNI 120 RAWWKEKVKF DKNGPAAIAK YEAECLAMCE ADNNRNGNSQ SILQDLQDAT LGSLLSSLMQ 180 HCEPPQRKFP LEKKVPPPWW PTGKEDWWVK LGLPHVQTPP YKKPHDLKKM WKVGVLTAVI 240 KHMSPDIAKI RRHVRQSKCL QDKMTAKESA IWLGVLSREE ALIRQPSSDN GMSGVTEIPP 300 GRGEKQVTAN SDSNYDVDGA DDGLGSVSSR DERRSQSMDV EPSGSQQNNT SQPGQDKEKP 360 DKPPKRKRPR LRSKPTDELP QPSQNEVMPI EPINNIPDIN YTEEVHLIGY QNDQNQQEDE 420 TVTGLRSLDK DIDVESQLPV SDFNSLYALP NNEVSTQSRH EGSVAAMHFH GIPNTGMHHS 480 DTYNFYNPSV DYEPKHERQR APIAMNESYV RPADGVHLPT VHRNSSEISG GDLPYYVNNE 540 QDRPVDTHFG SPLDSLSLDF GGFNSPLNFG IDGTSVLDDL LLDDDLIQYF GA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wij_A | 2e-72 | 160 | 291 | 9 | 140 | ETHYLENE-INSENSITIVE3-like 3 protein |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 365 | 371 | RKRPRLR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in the ethylene response pathway. {ECO:0000269|PubMed:9215635, ECO:0000269|PubMed:9851977}. | |||||
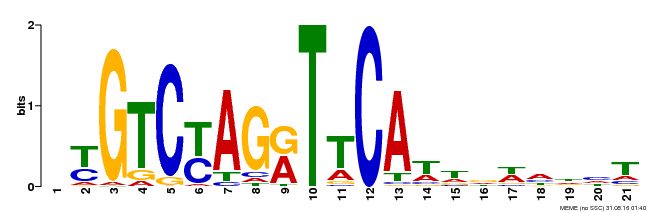
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00233 | DAP | Transfer from AT1G73730 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010091992.1 | 0.0 | ETHYLENE INSENSITIVE 3-like 3 protein | ||||
| Swissprot | O23116 | 0.0 | EIL3_ARATH; ETHYLENE INSENSITIVE 3-like 3 protein | ||||
| TrEMBL | A0A2P5EUP8 | 0.0 | A0A2P5EUP8_TREOI; Ethylene insensitive 3-like protein, DNA-binding domain containing protein | ||||
| STRING | XP_010091992.1 | 0.0 | (Morus notabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF322 | 34 | 200 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G73730.1 | 1e-169 | ETHYLENE-INSENSITIVE3-like 3 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



