 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PK00408.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Cannabaceae; Cannabis
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 727aa MW: 78004.5 Da PI: 6.1506 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 100.8 | 8.4e-32 | 299 | 355 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
+DgynWrKYGqK+vkgse+prsYY+Ct+++C+vkkkvers+ +++++ei+Y+g Hnh
PK00408.2 299 EDGYNWRKYGQKQVKGSEYPRSYYKCTHPNCQVKKKVERSH-EGHITEIIYKGAHNHA 355
7****************************************.***************7 PP
| |||||||
| 2 | WRKY | 102.8 | 1.9e-32 | 523 | 581 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
ldDgy+WrKYGqK+vkg+++prsYY+Ct agC+v+k+ver+++d k v++tYeg+Hnh+
PK00408.2 523 LDDGYRWRKYGQKVVKGNPNPRSYYKCTNAGCTVRKHVERASHDLKSVITTYEGKHNHD 581
59********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50811 | 22.986 | 293 | 357 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 7.06E-25 | 297 | 357 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 1.6E-27 | 297 | 357 | IPR003657 | WRKY domain |
| SMART | SM00774 | 1.8E-35 | 298 | 356 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 2.4E-24 | 299 | 354 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 9.2E-37 | 508 | 583 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.96E-28 | 515 | 583 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 38.32 | 518 | 583 | IPR003657 | WRKY domain |
| SMART | SM00774 | 1.9E-37 | 523 | 582 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 7.5E-25 | 524 | 581 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009555 | Biological Process | pollen development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0030010 | Biological Process | establishment of cell polarity | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 727 aa Download sequence Send to blast |
MAGLDDNVAI IGDWVPPSPS SRAFFSAMLG EDIGSRQFSG TTHNKTEDLF PNNNNNNNNN 60 FKEFREMGSF SSEQKSNSRG GGLVDRIAAR AGFNAPRLNT ESMRSSSSDL SLAASESRSP 120 YLTIPPGLSP TTLLDSPVFL SNSLAQPSPT TGKFPFASSN GNTRSSTLIM EASDRNKESF 180 FEDINAGSFA FKPVPNSGPS SSFFLGTATK MTSAAAMPPH PHPSYPNIEM SVQSENSPPQ 240 SVGGGNEKDS SSDQKGGELE LSPPLEEQGE EEGDQKGSGG DCSAGGGGGG GGGGGTISED 300 GYNWRKYGQK QVKGSEYPRS YYKCTHPNCQ VKKKVERSHE GHITEIIYKG AHNHAKPSHR 360 RSGIGSSSNP LADMRMDIQE QAGMQNGNVN VNCGGGDGDP VWASTQKGTV VDWRNDSLEV 420 TSSASASAGP EYCNQSTSLQ AQNGTQLESV DAVVDASSTF SNDEDEDDRG THGSASLGYE 480 GEEDESESKR RKLEAFATEM SGATRAIREP RVVVQTTSEV DILDDGYRWR KYGQKVVKGN 540 PNPRSYYKCT NAGCTVRKHV ERASHDLKSV ITTYEGKHNH DVPAARNSSH VNSGPSMQLQ 600 GSASTAMQSH VHRPEPSSQQ LHNNMGRLSS YSLPRQQLGN FSFGMNQAGL ANLAMAGLGS 660 MPPKLQGMPM NPYLAQQQQR QANEMSFMLP KGEPKAEPIS ESATGLNLSN ASVYQQFMSR 720 LPLGPQM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wj2_A | 7e-37 | 297 | 584 | 8 | 78 | Probable WRKY transcription factor 4 |
| 2lex_A | 7e-37 | 297 | 584 | 8 | 78 | Probable WRKY transcription factor 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Regulates WOX8 and WOX9 expression and basal cell division patterns during early embryogenesis. Interacts specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element. Required to repolarize the zygote from a transient symmetric state. {ECO:0000269|PubMed:21316593}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00071 | PBM | Transfer from AT5G56270 | Download |
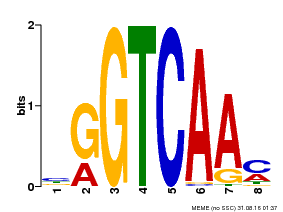
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_021826759.1 | 0.0 | probable WRKY transcription factor 2 | ||||
| Refseq | XP_021826760.1 | 0.0 | probable WRKY transcription factor 2 | ||||
| Refseq | XP_021826761.1 | 0.0 | probable WRKY transcription factor 2 | ||||
| Swissprot | Q9FG77 | 0.0 | WRKY2_ARATH; Probable WRKY transcription factor 2 | ||||
| TrEMBL | A0A314Y5Z5 | 0.0 | A0A314Y5Z5_PRUYE; Putative WRKY transcription factor 2 | ||||
| STRING | XP_008246415.1 | 0.0 | (Prunus mume) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF9524 | 34 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G56270.1 | 1e-169 | WRKY DNA-binding protein 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



