 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Carubv10026599m | ||||||||
| Common Name | CARUB_v10026599mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 378aa MW: 41699.5 Da PI: 4.7692 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 98.7 | 3.8e-31 | 189 | 247 | 2 | 59 |
--SS-EEEEEEE--TT-SS-EEEEEE-ST.T---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsa.gCpvkkkversaedpkvveitYegeHnhe 59
+D ++WrKYGqK++kgs++pr+YYrC+s+ gC ++k+vers+ dp+++++tY+geH+h+
Carubv10026599m 189 SDMWAWRKYGQKPIKGSPYPRNYYRCSSSkGCLARKQVERSNLDPNIFIVTYTGEHTHP 247
79*************************998****************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.20.25.80 | 4.5E-31 | 175 | 249 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 4.84E-27 | 181 | 249 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 33.281 | 183 | 249 | IPR003657 | WRKY domain |
| SMART | SM00774 | 6.1E-39 | 188 | 248 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 3.3E-26 | 190 | 247 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007263 | Biological Process | nitric oxide mediated signal transduction | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 378 aa Download sequence Send to blast |
MSSDDWDLNA VVRSCSSSVS TATTINTNPC IGHEEDDGGN CKQQQDPPLP PPSPLFTHIH 60 GDRETSSSCN ELQDSCKPFL PVTTTTTTTW SPPPLLPPSL ASSSAPNVLM KQEQVHHLES 120 SSSSQDHQKP PLGVRVFPPS TSSSSSVFVF RGQRDQLLQQ QSQPPLRSRK RKNQQKRTIC 180 HVTQENLSSD MWAWRKYGQK PIKGSPYPRN YYRCSSSKGC LARKQVERSN LDPNIFIVTY 240 TGEHTHPRPT HRNSLAGSTR NKSQHVNPIP KPDSSSGQTV LGSNTVKDEI HLSPTTPLKG 300 NDDVRGTNGD EDMMSQEVNM DEEEEEEEEE EVEEEDDEDD DVDDILIPNL AVRDRDDLFF 360 AGNFSSWSAG SAGDGGG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5w3x_B | 1e-19 | 190 | 250 | 18 | 80 | Disease resistance protein RRS1 |
| 5w3x_D | 1e-19 | 190 | 250 | 18 | 80 | Disease resistance protein RRS1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Interacts specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00561 | DAP | Transfer from AT5G52830 | Download |
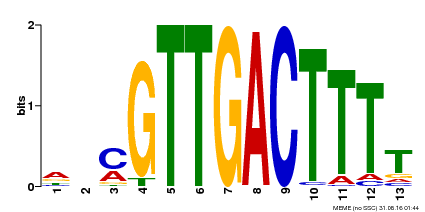
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Carubv10026599m |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006280638.1 | 0.0 | probable WRKY transcription factor 27 | ||||
| Swissprot | Q9FLX8 | 1e-147 | WRK27_ARATH; Probable WRKY transcription factor 27 | ||||
| TrEMBL | R0EXF5 | 0.0 | R0EXF5_9BRAS; Uncharacterized protein | ||||
| STRING | XP_006280638.1 | 0.0 | (Capsella rubella) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM9998 | 26 | 36 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52830.1 | 1e-122 | WRKY DNA-binding protein 27 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Carubv10026599m |
| Entrez Gene | 17874748 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



