 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Carubv10019974m | ||||||||
| Common Name | CARUB_v10019974mg | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 624aa MW: 71359.4 Da PI: 9.0023 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 95.5 | 4.9e-30 | 88 | 172 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW++ e+laL+++r+em++++r++ lk+plWee+s+km+e g++rs+k+Ckek+en+ k++k++keg+ ++++++ t+++f++lea
Carubv10019974m 88 RWPRPETLALLRIRSEMGKAFRDSTLKAPLWEEISRKMMELGYKRSAKKCKEKFENVYKYHKRTKEGRTGKSEGK--TYRFFEELEA 172
8********************************************************************975544..6******985 PP
| |||||||
| 2 | trihelix | 106.9 | 1.4e-33 | 442 | 527 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+k ev aLi++r+++e +++++ k+plWee+s+ mr+ g++r++k+Ckekwen+nk++kk+ke++kkr + +s+tcpyf+qlea
Carubv10019974m 442 RWPKTEVEALIRIRKNLEANYQENGTKGPLWEEISAGMRRLGYNRNAKRCKEKWENINKYFKKVKESNKKR-PLDSKTCPYFHQLEA 527
8*********************************************************************8.9************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00717 | 0.0046 | 85 | 147 | IPR001005 | SANT/Myb domain |
| CDD | cd12203 | 8.17E-24 | 87 | 152 | No hit | No description |
| PROSITE profile | PS50090 | 7.515 | 87 | 145 | IPR017877 | Myb-like domain |
| Pfam | PF13837 | 9.1E-20 | 87 | 172 | No hit | No description |
| PROSITE profile | PS50090 | 7.352 | 435 | 499 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 0.0056 | 439 | 501 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.6E-4 | 441 | 498 | IPR009057 | Homeodomain-like |
| Pfam | PF13837 | 2.6E-23 | 441 | 528 | No hit | No description |
| CDD | cd12203 | 1.96E-25 | 442 | 506 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 624 aa Download sequence Send to blast |
QNKKKKRKEK GKSTTGFVFS SNSPPHHISL SLFKKDPNSS FQVSMSGNSS GPLESSGGGA 60 GGSGEEEKDM KMMMEETGEV AGGGGGNRWP RPETLALLRI RSEMGKAFRD STLKAPLWEE 120 ISRKMMELGY KRSAKKCKEK FENVYKYHKR TKEGRTGKSE GKTYRFFEEL EAFETLNSYH 180 HPESQPAKSS ATLTTASLIP WISSNNNPST EKSSLPLKHH HHQVSVQPIT TNPTFLTKQP 240 SSTTPFPFYS NNNTTTLSQP PLSSDLMNNV SSLHLFSSST SSSTASDEEE DHHDQGKRSR 300 KRRRYWKGLF TKLTKELMDK QEKMQRRFLE TLENREKERI SREEAWRVQE IARINREHET 360 FLHERSNAAA KDAAIISFLH KISGGQQQQP QQQNHKPSQR KQYQSDHSIT FESKEPKTIL 420 LETTTKIGNY DTSHSISPSS SRWPKTEVEA LIRIRKNLEA NYQENGTKGP LWEEISAGMR 480 RLGYNRNAKR CKEKWENINK YFKKVKESNK KRPLDSKTCP YFHQLEALYN ERNKNGAMPL 540 PLPLPLMVTP ERQLLVSQET QTELETDQRD KVGDKEDEEE GESEEDEYDE EEEGEGDNET 600 SEFEIVLNKT SSSPMDINNN LFT* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 2 | 7 | KKKKRK |
| 2 | 130 | 138 | KRSAKKCKE |
| 3 | 299 | 303 | RKRRR |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds specific DNA sequence. {ECO:0000250}. | |||||
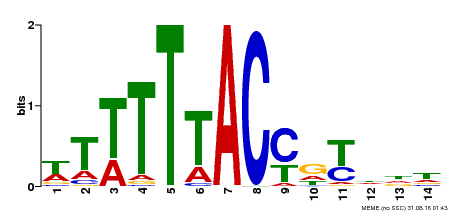
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00244 | DAP | Transfer from AT1G76890 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Carubv10019974m |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY087117 | 0.0 | AY087117.1 Arabidopsis thaliana clone 3190 mRNA, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006300884.2 | 0.0 | trihelix transcription factor GT-2 | ||||
| Swissprot | Q39117 | 0.0 | TGT2_ARATH; Trihelix transcription factor GT-2 | ||||
| TrEMBL | R0GDA8 | 0.0 | R0GDA8_9BRAS; Uncharacterized protein (Fragment) | ||||
| STRING | XP_006300884.1 | 0.0 | (Capsella rubella) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM5995 | 28 | 47 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G76890.2 | 0.0 | Trihelix family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Carubv10019974m |
| Entrez Gene | 17895445 |



