 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_9152_iso_9 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 730aa MW: 79327.1 Da PI: 6.4782 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 101.9 | 3.7e-32 | 307 | 364 | 2 | 60 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS-- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhek 60
+DgynWrKYGqK+vkgse+prsYY+Ct+++C+vkkkvers+ +++++ei+Y+g Hnh+k
cra_locus_9152_iso_9_len_2518_ver_3 307 EDGYNWRKYGQKQVKGSEYPRSYYKCTHQNCQVKKKVERSQ-EGHITEIIYKGAHNHPK 364
8****************************************.***************85 PP
| |||||||
| 2 | WRKY | 103.7 | 1e-32 | 519 | 577 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
ldDgy+WrKYGqK+vkg+++prsYY+CtsagC+v+k+ver+++d k v++tYeg+Hnh+
cra_locus_9152_iso_9_len_2518_ver_3 519 LDDGYRWRKYGQKVVKGNPNPRSYYKCTSAGCTVRKHVERASHDLKSVITTYEGKHNHD 577
59********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.20.25.80 | 1.2E-28 | 304 | 365 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.28E-25 | 305 | 365 | IPR003657 | WRKY domain |
| SMART | SM00774 | 4.4E-36 | 306 | 364 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 9.0E-25 | 307 | 363 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 23.365 | 307 | 365 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 5.4E-37 | 504 | 579 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.7E-28 | 511 | 579 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 38.664 | 514 | 579 | IPR003657 | WRKY domain |
| SMART | SM00774 | 1.5E-37 | 519 | 578 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 6.6E-25 | 520 | 577 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009555 | Biological Process | pollen development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0030010 | Biological Process | establishment of cell polarity | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 730 aa Download sequence Send to blast |
MGDWMPPSPS PRTLLSMLGD DVGSRSISDP IGEKIEGFSF PGPQEYMAST NADEKNGSQA 60 GLGADEAAKF SGMYEQKMSS RGGLLERRAA RAGFNAPRLN TESIRAADLL QNQEVRSPYL 120 TIPPGLSPTT LLDSPVFVNN SLVQPSPTTG KFQFVSNANS RSSTVMTEFP DKSRETFFGN 180 INPSSFAFRP MAESGPSLFF GTSNRVNPTS LPQQSFPSIE VSVQSQNVHP SSLERPNTTA 240 LNQQADFSKP STEKETGVSN NNIASEPRTL YNVGGGMDHS PPLDEQQDED ADQRSNGDPN 300 IGGAPAEDGY NWRKYGQKQV KGSEYPRSYY KCTHQNCQVK KKVERSQEGH ITEIIYKGAH 360 NHPKPPPNRR SALGSSNALS DMQLENSEQA GTGADGDPLW ANVQRGNGAS EGRNDNPEVT 420 SSAMMGSEYC NGANSLQAQN GSQFETGDAV DASSTFSNDE EDDDRATHGS VSLGYDGEGD 480 ESESKRRKIE TYPTDMSGAS RAIREPRVVV QTTSEVDILD DGYRWRKYGQ KVVKGNPNPR 540 SYYKCTSAGC TVRKHVERAS HDLKSVITTY EGKHNHDVPA ARNSSHVNSG GASCTLPTQP 600 AAVQTQVHRP EPSSQLHNSM TQFERPSSLA SFALPGRPPL GPPNHHHGFG FGINQHGLSN 660 LGLTGLGHNQ IRIPSLPVHQ YLGQQPRQPM NDMGLMLPKG EPKVEPVSDP GLNHSNGSSV 720 YHQVMHRMQM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wj2_A | 1e-36 | 307 | 580 | 8 | 78 | Probable WRKY transcription factor 4 |
| 2lex_A | 1e-36 | 307 | 580 | 8 | 78 | Probable WRKY transcription factor 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Regulates WOX8 and WOX9 expression and basal cell division patterns during early embryogenesis. Interacts specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element. Required to repolarize the zygote from a transient symmetric state. {ECO:0000269|PubMed:21316593}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00071 | PBM | Transfer from AT5G56270 | Download |
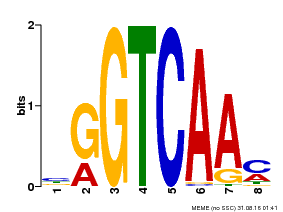
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027170840.1 | 0.0 | probable WRKY transcription factor 2 isoform X1 | ||||
| Refseq | XP_027170841.1 | 0.0 | probable WRKY transcription factor 2 isoform X1 | ||||
| Refseq | XP_027170842.1 | 0.0 | probable WRKY transcription factor 2 isoform X1 | ||||
| Refseq | XP_027170844.1 | 0.0 | probable WRKY transcription factor 2 isoform X1 | ||||
| Refseq | XP_027170845.1 | 0.0 | probable WRKY transcription factor 2 isoform X1 | ||||
| Swissprot | Q9FG77 | 0.0 | WRKY2_ARATH; Probable WRKY transcription factor 2 | ||||
| TrEMBL | A0A336RDJ4 | 0.0 | A0A336RDJ4_CATRO; WRKY transcription factor 2 | ||||
| STRING | VIT_19s0015g01870.t01 | 0.0 | (Vitis vinifera) | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



