 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_625_iso_6 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 464aa MW: 50055.8 Da PI: 7.3101 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 31.1 | 4.1e-10 | 296 | 344 | 3 | 55 |
HHHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 3 rahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+h+ +Er RR+ri +++ L+el+P + +k Ka +L + ++Y++sLq
cra_locus_625_iso_6_len_2866_ver_3 296 NSHSLAERVRRERISERMRLLQELVPGC----NKITGKAVMLDEIINYVQSLQ 344
58**************************....999*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 7.85E-17 | 291 | 360 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 3.03E-11 | 291 | 348 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 3.4E-16 | 293 | 361 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 15.472 | 293 | 343 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.6E-7 | 296 | 344 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.1E-10 | 299 | 349 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009637 | Biological Process | response to blue light | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 464 aa Download sequence Send to blast |
MATKDNVGFQ PTGGGNNILN CPSSGMDSNL ISDKVAMCSD SMFKNSSTGV ETFYGSGWDP 60 IVSLNHQENY GGSSVVPHNE FANSDYPVVL GNQSMGSSSH LVHYPSDSGL VDMLPKISCF 120 GSGSFSEMVS SFGLQDCGQV TETGSHINGT SNKGVAYQEG GQNSGGGALG SSSNGKKKRK 180 TSDSPSPLHS KKVKLPDFLS SYNFGVLFQN VIFQRVIYII ICLLKNVEGE QQKDLSGNSS 240 ECSKEQDEMK QIMEQNNSTN FRSKQAGKQI KDNSSNGEPS KETYIHVRAK RGQATNSHSL 300 AERVRRERIS ERMRLLQELV PGCNKITGKA VMLDEIINYV QSLQQQVEFL SMKLATVNPE 360 LNIDLDQIFS KDILHASRGT SAAATLGIRP TMNPSHAFPG FPPGTFPGIP GATPPFHPPL 420 PQAVWDSELQ NVFAMGFDSN PSANNLGPNA GRSKLDLXRV GVKA |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00135 | DAP | Transfer from AT1G10120 | Download |
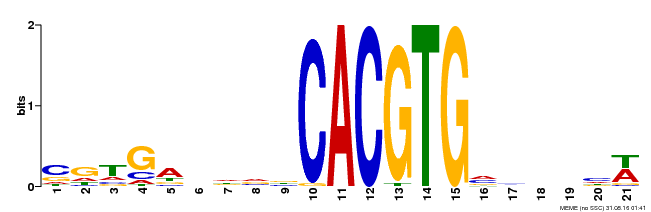
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027092757.1 | 0.0 | transcription factor bHLH74-like isoform X1 | ||||
| Refseq | XP_027092758.1 | 0.0 | transcription factor bHLH74-like isoform X1 | ||||
| TrEMBL | A0A438KMH0 | 1e-165 | A0A438KMH0_VITVI; Transcription factor bHLH74 | ||||
| STRING | VIT_18s0001g08600.t01 | 1e-164 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA5250 | 24 | 35 |



