 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_4684_iso_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 479aa MW: 52073.4 Da PI: 8.2476 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 27.9 | 5.4e-09 | 56 | 83 | 1 | 29 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIar 29
r rWT+eE++++++a k++G W++I
cra_locus_4684_iso_1_len_2600_ver_3 56 RERWTEEEHKKFLEALKLYGRA-WRRIEG 83
78******************88.***965 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.82E-7 | 41 | 82 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.5E-7 | 54 | 82 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 2.0E-5 | 56 | 83 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 5.1E-6 | 56 | 81 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.63E-4 | 58 | 81 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 479 aa Download sequence Send to blast |
MAIQGQSNNS ESSSTISIGN QISLHVDIPS TKNEQFQCED DCLPKVRKPY TITKQRERWT 60 EEEHKKFLEA LKLYGRAWRR IEGCAESTNG DSGSGKVIEI PPPRPKRKPL HPYPRKLVSP 120 AKSGTATSQK LTQTVSPNIS SPAEEHQSPT SVLSAPCSDT PGTTDSATSD GSESLSPISS 180 VVGAKSVGFV LSELPDLTSE TKRSPSSSQV NSSADNKSPS TSQVNNCANQ ADQLHLKLEL 240 FPQDNAFDKE GSVEVSSSQI FKLFGKTVLV IDPSRPSSST PGAGKVQPSH QNDGKSNSTK 300 VPLGESECLR RALPCKARGN LYPSSLQREC PDTLLCGPFH SLPWLMMYQG DSSASFVQVH 360 SPIAIKAQPL LDKKELHRVQ SCKGGSSTGS DAESESIETV EDKNIEVKFR ELSQEGEVQK 420 QKLPLLKPGA KAASFFRQKA NSSKCGKGFV PYKRCLSDRD TLSAITGEER EEQRVRLCL |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00515 | DAP | Transfer from AT5G17300 | Download |
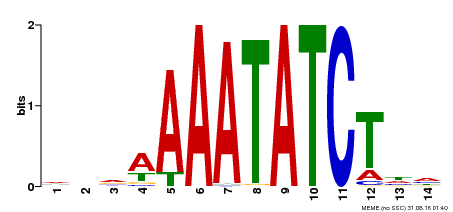
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027075006.1 | 1e-125 | protein REVEILLE 1-like isoform X1 | ||||
| TrEMBL | A1DR85 | 0.0 | A1DR85_CATRO; MYB transcription factor (Fragment) | ||||



