 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_4179_iso_3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 421aa MW: 45978.6 Da PI: 5.2736 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 36.4 | 9.4e-12 | 364 | 409 | 6 | 55 |
HHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 6 nerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
+++Er RR ri +++ +L+e +P+ +k + +++L +Av+YIk+Lq
cra_locus_4179_iso_3_len_1742_ver_3 364 SIAERVRRTRISERMRKLQEAVPNM----DKQTNTSDMLDLAVDYIKDLQ 409
689*********************7....688*****************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50888 | 14.297 | 358 | 408 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 1.14E-14 | 361 | 417 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 3.99E-12 | 362 | 413 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 3.7E-13 | 363 | 417 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 2.8E-12 | 364 | 414 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 9.8E-9 | 364 | 409 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 421 aa Download sequence Send to blast |
MTGGLTRYRS APSSYFTSFL DATDNYVSSS VGGGGGGIGG MGGYVMDDLD QFLNRFMPNN 60 SGQRQEELNP GNNFSNSMSI QSHQTQFVGS MKQEQESMES QGQLNDFSSQ MMYQSQDSAE 120 IQAPAQRNQQ NSEASVDYSN RLLNSVNTNR FTPMKMATGG GNSNLIRYNS SPAGLFANIN 180 IENEYGAMRG MGNFGAGNNA NAEASFSSTS RFEGFSSGQS SSGLMPPISE MGTKGMGDQG 240 TSFEGQRNDG GGNYITGFPV SSWDDSGMLS ENLGKGLVGD NNNQQVRFYA LYAIVFRVXC 300 PSDFVYLFGN QGNEGQKDPA PTLLSRHLSL PSTSVEFSAM ETLMQDSVLC KIRAKRGCAT 360 HPRSIAERVR RTRISERMRK LQEAVPNMDK QTNTSDMLDL AVDYIKDLQK QLKTLLDNRA 420 K |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00646 | PBM | Transfer from LOC_Os08g39630 | Download |
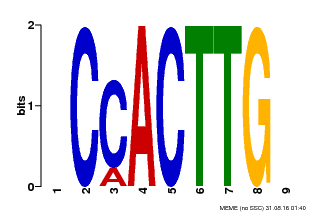
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027077343.1 | 1e-155 | transcription factor bHLH130-like isoform X1 | ||||
| Refseq | XP_027077352.1 | 1e-155 | transcription factor bHLH130-like isoform X1 | ||||
| Refseq | XP_027077359.1 | 1e-155 | transcription factor bHLH130-like isoform X1 | ||||
| TrEMBL | A0A068VCJ1 | 1e-151 | A0A068VCJ1_COFCA; Uncharacterized protein | ||||
| STRING | PGSC0003DMT400061284 | 1e-104 | (Solanum tuberosum) | ||||



