 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_384_iso_2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 269aa MW: 31129.9 Da PI: 8.3539 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 22.4 | 3.3e-07 | 107 | 128 | 2 | 23 |
EETTTTEEESSHHHHHHHHHHT CS
zf-C2H2 2 kCpdCgksFsrksnLkrHirtH 23
+C+ Cg sF+ + +L++H+++H
cra_locus_384_iso_2_len_938_ver_3 107 VCHECGVSFKKPAYLRQHMQSH 128
6*******************99 PP
| |||||||
| 2 | zf-C2H2 | 18.7 | 4.9e-06 | 160 | 184 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
++Cp dC+ s++rk++L+rH+ H
cra_locus_384_iso_2_len_938_ver_3 160 FTCPvdDCHSSYRRKDHLTRHLLQH 184
89*******************9887 PP
| |||||||
| 3 | zf-C2H2 | 18.3 | 6.5e-06 | 189 | 214 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
++Cp +C+++F + n++rH++ H
cra_locus_384_iso_2_len_938_ver_3 189 FECPvdNCKRRFAFQGNMRRHMKEfH 214
89*********************988 PP
| |||||||
| 4 | zf-C2H2 | 20.4 | 1.3e-06 | 228 | 252 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirtH 23
y+C+ Cgk+F+ s L++H +H
cra_locus_384_iso_2_len_938_ver_3 228 YVCSeiGCGKVFKYASKLRKHEDSH 252
99*******************9888 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF57667 | 8.26E-7 | 60 | 129 | No hit | No description |
| SMART | SM00355 | 32 | 62 | 84 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 4.5E-6 | 98 | 129 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0098 | 106 | 128 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.658 | 106 | 133 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 108 | 128 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 8.1E-9 | 157 | 186 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.0059 | 160 | 184 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 9.64 | 160 | 189 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 162 | 184 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 6.16E-7 | 170 | 216 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 1.7E-6 | 187 | 211 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 0.005 | 189 | 214 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 10.221 | 189 | 219 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 191 | 214 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 1.5E-7 | 225 | 252 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SuperFamily | SSF57667 | 4.05E-5 | 225 | 252 | No hit | No description |
| SMART | SM00355 | 0.0015 | 228 | 252 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 11.053 | 228 | 252 | IPR007087 | Zinc finger, C2H2 |
| PROSITE pattern | PS00028 | 0 | 230 | 252 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008097 | Molecular Function | 5S rRNA binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| GO:0080084 | Molecular Function | 5S rDNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 269 aa Download sequence Send to blast |
XXWELAALAE CECALLSTPG FLTKSGAPHR NSSTSPLSEI VSKVEMVEEK SKPVIFRDIR 60 RYYCEYCGIC RSKKALISAH ILSHHQDDLK QGEEDKRETH NGPKMNVCHE CGVSFKKPAY 120 LRQHMQSHSL EAVKHWIPHK ELQNARGQCX LECDQYQRPF TCPVDDCHSS YRRKDHLTRH 180 LLQHQGKLFE CPVDNCKRRF AFQGNMRRHM KEFHMEPSSS DEVSAKQYVC SEIGCGKVFK 240 YASKLRKHED SHGIILSILC WLVLQSFVQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1tf6_A | 1e-11 | 157 | 260 | 40 | 137 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| 1tf6_D | 1e-11 | 157 | 260 | 40 | 137 | PROTEIN (TRANSCRIPTION FACTOR IIIA) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Essential protein (PubMed:22353599). Isoform 1 is a transcription activator the binds both 5S rDNA and 5S rRNA and stimulates the transcription of 5S rRNA gene (PubMed:12711688, PubMed:22353599). Isoform 1 regulates 5S rRNA levels during development (PubMed:22353599). {ECO:0000269|PubMed:12711688, ECO:0000269|PubMed:22353599}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
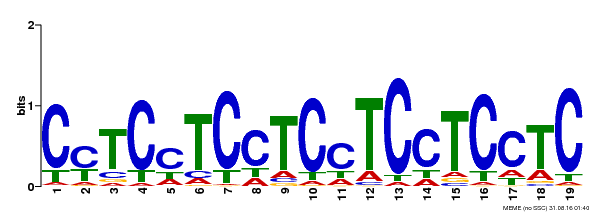
| Motif ID | Method | Source | Motif file |
| MP00229 | DAP | Transfer from AT1G72050 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002278827.1 | 8e-86 | PREDICTED: transcription factor IIIA | ||||
| Swissprot | Q84MZ4 | 2e-42 | TF3A_ARATH; Transcription factor IIIA | ||||
| TrEMBL | D7TQ66 | 3e-89 | D7TQ66_VITVI; Uncharacterized protein | ||||
| STRING | VIT_08s0040g00240.t01 | 5e-90 | (Vitis vinifera) | ||||



