 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_3425_iso_2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 313aa MW: 32926.4 Da PI: 9.7532 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 119.8 | 3.3e-37 | 55 | 162 | 2 | 89 |
TCP 2 agkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssasecea 75
a+k+++++++hTkv+gR+RR+R++a+caar+F+L++eLG+++d++ti+WLlqqa+p+i+++tgt++++as+ a
cra_locus_3425_iso_2_len_1968_ver_3 55 APKRSSNKDRHTKVEGRGRRIRMPALCAARIFQLTRELGHKSDGETIQWLLQQAEPSIIAATGTGTIPASALAA 128
789******************************************************************77744 PP
TCP 76 esssssasnsssg....................k 89
+ s s++ ss++
cra_locus_3425_iso_2_len_1968_ver_3 129 AGGSFSQHDSSISaglhqkieelggahmgdsggG 162
4444444444444344333335555444444441 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 1.7E-29 | 60 | 156 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 28.016 | 61 | 115 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008361 | Biological Process | regulation of cell size | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000987 | Molecular Function | core promoter proximal region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 313 aa Download sequence Send to blast |
MDIDPKGSKQ PQQEVPRSFL NNMGENKKSA EMKDFQISIA DKKEDNNSSK KQQLAPKRSS 60 NKDRHTKVEG RGRRIRMPAL CAARIFQLTR ELGHKSDGET IQWLLQQAEP SIIAATGTGT 120 IPASALAAAG GSFSQHDSSI SAGLHQKIEE LGGAHMGDSG GGNSRTSNWP PSASAGGNFG 180 SRPHMATAAG IWPPSAAAFS GFGFPQPSSS SSSSGPSSTN LGSENANYLQ KIGFSGFDLP 240 NSNLGSMSFT SILGGAPGLE LGLAQDGHIG VLNPQTLSQI YQQMETAKMH HQQQQQQKPP 300 SSKDNSQGSG GGQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds to the site II motif (3'-TGGGCC/T-5') in the promoter of PCNA-2 and to 3'-GCCCG/A-5' elements in the promoters of cyclin CYCB1-1 and ribosomal protein genes. {ECO:0000269|PubMed:12631321, ECO:0000269|PubMed:16123132}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00636 | PBM | Transfer from Glyma.19G095300 | Download |
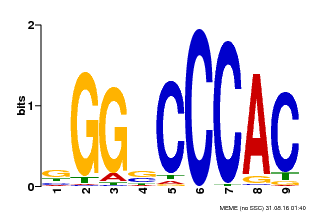
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022750458.1 | 1e-124 | transcription factor TCP20-like | ||||
| Refseq | XP_022750459.1 | 1e-124 | transcription factor TCP20-like | ||||
| Swissprot | Q9LSD5 | 1e-74 | TCP20_ARATH; Transcription factor TCP20 | ||||
| TrEMBL | V7CWP0 | 1e-122 | V7CWP0_PHAVU; Uncharacterized protein | ||||
| STRING | XP_007161676.1 | 1e-123 | (Phaseolus vulgaris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA248 | 24 | 201 |



