 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_15469_iso_2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 442aa MW: 45688.8 Da PI: 6.9623 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 37 | 6.1e-12 | 229 | 274 | 5 | 55 |
HHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 5 hnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
h+++Er RR+ri +++ L+el+P+a K +Ka++L + ++Y+k Lq
cra_locus_15469_iso_2_len_1875_ver_3 229 HSIAERLRRERIAERMKALQELVPNA-----NKTDKASMLDEIIDYVKFLQ 274
99***********************8.....6******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 5.23E-17 | 222 | 284 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 3.06E-13 | 222 | 278 | No hit | No description |
| PROSITE profile | PS50888 | 17.39 | 224 | 273 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.8E-16 | 226 | 283 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.0E-9 | 229 | 274 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.5E-13 | 230 | 279 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048767 | Biological Process | root hair elongation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 442 aa Download sequence Send to blast |
MTLHDLQNSH QQQNHFDPSN SQDDFLEQIL ISSSIPSSWP PDLSASATNS GSKPQSPWDS 60 LPQNPLASLE DQSALLTSKL RQHQIGGAKA MMLQQQLMLS RGLAGNGLRS PTTAAGNGGL 120 LPMPLSLNGN GGDVIDGSPF KSANPANDAS VQALYNGFAV SLGQTSNQSQ HFHHSQGGAM 180 QAQNFGATAG GAAPTMTQTT ASGSAGTGAA AQPRQQRVRA RRGQATDPHS IAERLRRERI 240 AERMKALQEL VPNANKTDKA SMLDEIIDYV KFLQLQVKVL SMSRLGGAAA VAPLVADMSS 300 EGGGDCVQGN GRNGNGGTQA ASSSNNDSMT VTEHQVAKLM EEDMGSAMQY LQGKGLCLMP 360 ISLATAISTA TCRNPMTSTN HHAHNGNPLL SSSAVNGGGI NGGEGGAAPS SPSMSVLTVQ 420 SATIGNVGAD MSVKDATSVS KP |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 233 | 238 | RLRRER |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00659 | PBM | Transfer from Pp3c14_15890 | Download |
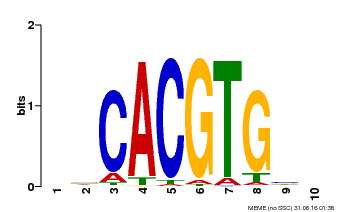
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By cold, UV, ethylene (ACC), flagellin, jasmonic acid (JA), and salicylic acid (SA) treatments. {ECO:0000269|PubMed:12679534}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027072432.1 | 0.0 | transcription factor bHLH66-like | ||||
| Swissprot | Q9ZUG9 | 4e-86 | BH066_ARATH; Transcription factor bHLH66 | ||||
| TrEMBL | A0A068UPR8 | 1e-179 | A0A068UPR8_COFCA; Uncharacterized protein | ||||
| STRING | VIT_11s0037g00040.t01 | 1e-159 | (Vitis vinifera) | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



