 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_15147_iso_5 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 276aa MW: 31259.3 Da PI: 6.2627 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 37.7 | 4.6e-12 | 33 | 63 | 4 | 34 |
XCHHHCHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 4 lkrerrkqkNReAArrsRqRKkaeieeLeek 34
+k+ rr+++NReAAr+sR+RKka++++Le+
cra_locus_15147_iso_5_len_827_ver_3 33 DKVLRRLAQNREAARKSRLRKKAYVQQLENS 63
7999*************************85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 1.7E-8 | 27 | 76 | No hit | No description |
| SMART | SM00338 | 2.0E-6 | 30 | 90 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 8.749 | 32 | 76 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 9.8E-8 | 33 | 63 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 3.13E-7 | 34 | 76 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 37 | 52 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 2.0E-29 | 121 | 195 | IPR025422 | Transcription factor TGA like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 276 aa Download sequence Send to blast |
VEAKLDNQSE DTSQGTYGPP SGKYDQEASK PIDKVLRRLA QNREAARKSR LRKKAYVQQL 60 ENSKLRLMQL EQELERARQQ GAYVGSGLDI SQISSSGIGH SGNTAFGIEY GNWVEERNRR 120 INDLKNALDS NISDGELHLL VDGAMNHYYD LFRLKALAAK SDVFYLLSGM WKTSVERLFL 180 WMGGFRPSEL LQIIMTPINP LTDEQLLHIY NLRQSCQQAE DALTQGMEKL HQYLGEAVLA 240 GQLGQGNYLP QMEAAMEKLE ALVRFVNQAD HLRQET |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-TGACG-3'. Recognizes ocs elements like the as-1 motif of the cauliflower mosaic virus 35S promoter. Binding to the as-1-like cis elements mediate auxin- and salicylic acid-inducible transcription. Could also bind to the Hex-motif (5'-TGACGTGG-3') another cis-acting element found in plant histone promoters. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00642 | SELEX | Transfer from LOC107771743 | Download |
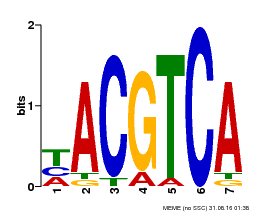
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027077188.1 | 1e-142 | TGACG-sequence-specific DNA-binding protein TGA-1A-like | ||||
| Refseq | XP_027179172.1 | 1e-142 | TGACG-sequence-specific DNA-binding protein TGA-1A-like | ||||
| Swissprot | P14232 | 1e-134 | TGA1A_TOBAC; TGACG-sequence-specific DNA-binding protein TGA-1A | ||||
| TrEMBL | A0A068TYL9 | 1e-141 | A0A068TYL9_COFCA; Uncharacterized protein | ||||
| STRING | VIT_07s0031g01320.t01 | 1e-141 | (Vitis vinifera) | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



