 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_13768_iso_1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 294aa MW: 31882.7 Da PI: 7.9491 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 29.3 | 2.1e-09 | 8 | 55 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT....-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg....tWktIartmgkgRtlkqcksrwqky 47
+W+ eE++ + +a++++ + +W++I+++++ ++ ++k ++q++
cra_locus_13768_iso_1_len_1174_ver_3 8 SWSREEEKAFENAIAMHWIEdskeQWEKISSMVP-SKSIDELKQHYQML 55
7****************99999************.************86 PP
| |||||||
| 2 | Myb_DNA-binding | 38.3 | 3.1e-12 | 134 | 178 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ l++ + ++G+g+W++I+r Rt+ q+ s+ qky
cra_locus_13768_iso_1_len_1174_ver_3 134 PWTEEEHRLFLLGLDKFGKGDWRSISRNYVISRTPTQVASHAQKY 178
7*****************************89************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 9.518 | 4 | 60 | IPR017884 | SANT domain |
| SMART | SM00717 | 4.5E-6 | 5 | 58 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.2E-4 | 7 | 55 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.1E-9 | 8 | 62 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 3.1E-7 | 8 | 55 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.62E-8 | 8 | 56 | No hit | No description |
| PROSITE profile | PS51294 | 16.887 | 127 | 183 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 8.61E-17 | 129 | 184 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.7E-10 | 131 | 181 | IPR001005 | SANT/Myb domain |
| TIGRFAMs | TIGR01557 | 3.2E-17 | 131 | 181 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 2.0E-10 | 132 | 177 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 9.6E-11 | 134 | 178 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.30E-9 | 134 | 179 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 294 aa Download sequence Send to blast |
MAASYSVSWS REEEKAFENA IAMHWIEDSK EQWEKISSMV PSKSIDELKQ HYQMLVEDVG 60 AIEAGNVPIP NYIGEEASSS SKDQQQAFSG IAATEKRSNC GYGGGFSGLG HDSSSAQGGK 120 GSSRAEQERR KGIPWTEEEH RLFLLGLDKF GKGDWRSISR NYVISRTPTQ VASHAQKYFI 180 RLNSMNRDRR RSSIHDITSV NNGDVSAHQA PITGQQNVPS TANPAALGPA MKHRGQPGMH 240 GLGVYGAPVG HPVVAPPGHI ASAVGTPVML PPGHHPPYVL PMAYPMAPPP TMHQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to 5'-TATCCA-3' elements in gene promoters. Derepresses strongly the sugar-repressed transcription of promoters containing SRS or 5'-TATCCA-3' elements. Functions with GAMYB to integrate diverse nutrient starvation and gibberellin (GA) signaling pathways during germination of grains. Sugar, nitrogen and phosphate starvation signals converge and interconnect with GA to promote the co-nuclear import of MYBS1 and GAMYB, resulting in the expression of a large set of GA-inducible hydrolases, transporters, and regulators that are essential for mobilization of nutrient reserves in the endosperm to support seedling growth (PubMed:22773748). {ECO:0000269|PubMed:12172034, ECO:0000269|PubMed:22773748}. | |||||
| UniProt | Transcription activator that binds to 5'-TATCCA-3' elements in gene promoters. Derepress strongly the sugar-repressed transcription of promoters containing SRS or 5'-TATCCA-3' elements. {ECO:0000250|UniProtKB:Q8LH59}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00191 | DAP | Transfer from AT1G49010 | Download |
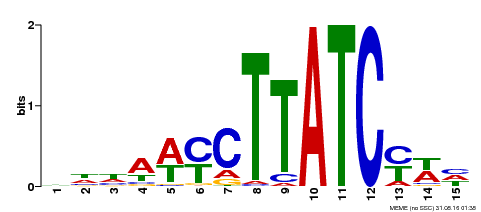
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by sucrose and gibberellic acid (GA). {ECO:0000269|PubMed:12172034}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018825411.1 | 1e-163 | PREDICTED: transcription factor DIVARICATA-like | ||||
| Swissprot | B8A9B2 | 5e-83 | MYBS1_ORYSI; Transcription factor MYBS1 | ||||
| Swissprot | Q8LH59 | 5e-83 | MYBS1_ORYSJ; Transcription factor MYBS1 | ||||
| TrEMBL | A0A2I4F186 | 1e-162 | A0A2I4F186_JUGRE; transcription factor DIVARICATA-like | ||||
| STRING | PGSC0003DMT400014350 | 1e-159 | (Solanum tuberosum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA5801 | 21 | 36 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



