 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | cra_locus_12712_iso_3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Gentianales; Apocynaceae; Rauvolfioideae; Vinceae; Catharanthinae; Catharanthus
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 339aa MW: 38427.8 Da PI: 8.7522 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 164.1 | 5.1e-51 | 26 | 153 | 3 | 128 |
NAM 3 pGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpkkvkaeekewyfFskrdkkyatgkrkn 75
pGfrFhPtdeelv +yL++kve++++++ e ik++diyk++Pw+Lpk+ + ++kewyfF+kr +ky+++ r+n
cra_locus_12712_iso_3_len_1166_ver_3 26 PGFRFHPTDEELVGFYLRRKVEKRPISI-ELIKQIDIYKHDPWNLPKASNVGDKEWYFFCKRGRKYRNSIRPN 97
9***************************.89***************98889********************** PP
NAM 76 ratksgyWkatgkdkevlsk...kgelvglkktLvfykgrapkgektdWvmheyrl 128
r+t sg+Wkatg d++++s+ ++ +glkk+Lv+y+g+a kg+ktdW+mhe+rl
cra_locus_12712_iso_3_len_1166_ver_3 98 RVTGSGFWKATGIDRPIYSAggeGRDCIGLKKSLVYYRGSAGKGTKTDWMMHEFRL 153
*******************96555555***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51005 | 53.535 | 24 | 185 | IPR003441 | NAC domain |
| SuperFamily | SSF101941 | 7.06E-55 | 25 | 185 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.1E-25 | 26 | 153 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005992 | Biological Process | trehalose biosynthetic process | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006561 | Biological Process | proline biosynthetic process | ||||
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process | ||||
| GO:0010120 | Biological Process | camalexin biosynthetic process | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 339 aa Download sequence Send to blast |
XRVEDMNGGG VIVSGGKDEE DDVPLPGFRF HPTDEELVGF YLRRKVEKRP ISIELIKQID 60 IYKHDPWNLP KASNVGDKEW YFFCKRGRKY RNSIRPNRVT GSGFWKATGI DRPIYSAGGE 120 GRDCIGLKKS LVYYRGSAGK GTKTDWMMHE FRLPADHDLK TTKHIDAKTI AQEAEVWTLC 180 RIFKRNVSYR KCMPEWKDHQ PSSAKKLNPN NNNNNINTDA SSKACSFESN NDHDIRQTYI 240 SFSTSSPVIT TNNNQYDERK PLLFSGHADH ITRQMFMGQS HHHHHHQAPI SSNATLMSSA 300 CSEVTDQFFK HGDWDELRSV VEFGAAAAAA SAGSPSFLL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 1e-48 | 26 | 191 | 19 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 1e-48 | 26 | 191 | 19 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 1e-48 | 26 | 191 | 19 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 1e-48 | 26 | 191 | 19 | 171 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 1e-48 | 26 | 191 | 22 | 174 | NAC domain-containing protein 19 |
| 3swm_B | 1e-48 | 26 | 191 | 22 | 174 | NAC domain-containing protein 19 |
| 3swm_C | 1e-48 | 26 | 191 | 22 | 174 | NAC domain-containing protein 19 |
| 3swm_D | 1e-48 | 26 | 191 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_A | 1e-48 | 26 | 191 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_B | 1e-48 | 26 | 191 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_C | 1e-48 | 26 | 191 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_D | 1e-48 | 26 | 191 | 22 | 174 | NAC domain-containing protein 19 |
| 4dul_A | 1e-48 | 26 | 191 | 19 | 171 | NAC domain-containing protein 19 |
| 4dul_B | 1e-48 | 26 | 191 | 19 | 171 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds to the 5'- RRYGCCGT-3' consensus core sequence. Central longevity regulator. Negative regulator of leaf senescence. Modulates cellular H(2)O(2) levels and enhances tolerance to various abiotic stresses through the regulation of DREB2A. {ECO:0000269|PubMed:22345491}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00310 | DAP | Transfer from AT2G43000 | Download |
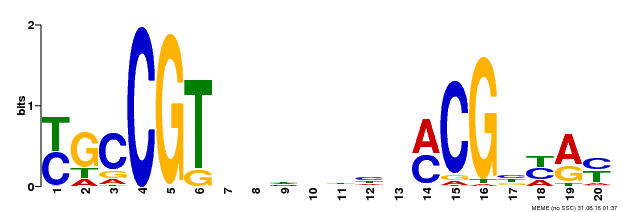
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by H(2)O(2), paraquat, ozone, 3-aminotriazole and salt stress. {ECO:0000269|PubMed:22345491}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_027061099.1 | 1e-157 | transcription factor JUNGBRUNNEN 1-like | ||||
| Refseq | XP_027172552.1 | 1e-157 | transcription factor JUNGBRUNNEN 1 | ||||
| Swissprot | Q9SK55 | 3e-93 | NAC42_ARATH; Transcription factor JUNGBRUNNEN 1 | ||||
| TrEMBL | A0A068UZ64 | 1e-147 | A0A068UZ64_COFCA; Uncharacterized protein | ||||
| STRING | XP_009591286.1 | 1e-130 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1697 | 24 | 69 |



