 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cre12.g514400.t2.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Chlorophyceae; Chlamydomonadales; Chlamydomonadaceae; Chlamydomonas
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 3234aa MW: 309339 Da PI: 7.5327 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 50 | 7e-16 | 50 | 94 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT +E++++v+a k++G W++I +++g ++t+ q++s+ qky
Cre12.g514400.t2.1 50 RERWTDDEHQRFVEALKLYGRA-WRKIEEYVG-TKTAVQIRSHAQKY 94
78******************88.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 7.33E-16 | 44 | 100 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 23.224 | 45 | 99 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 7.1E-8 | 45 | 98 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 6.7E-17 | 48 | 97 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 4.4E-13 | 49 | 97 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 3.6E-13 | 50 | 94 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.89E-8 | 52 | 95 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010243 | Biological Process | response to organonitrogen compound | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043496 | Biological Process | regulation of protein homodimerization activity | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 3234 aa Download sequence Send to blast |
MAHQFNGPGG PGSSVSPPPQ LKQEAVPAAQ NAAEPPPSKT RKQYTVSKRR ERWTDDEHQR 60 FVEALKLYGR AWRKIEEYVG TKTAVQIRSH AQKYFNKLEK AGDAAEIQVP PPRPKRKSVT 120 AAQRAAAAAA AAEQQLNPRS GSEERGYGDL QDDNDAGSEG GAEDGEPGYG QHDQMGSPDD 180 VPLAADLQGL LLQPPRNGAA AAAAAAAAAA AAGDLGPLAA AAGLPQLPVL QQQQQEQSAV 240 MQQQLQELQQ LQLQPFTAPG LMPFRGAAGA AAVQLPVLAA AAAAGQLLAA AAAGALALPP 300 APSPPPVLLG GALPPVGVGA LGQQPQQQPQ QATPSGTTLS AQAPMQMQAS PQQQQSQQQQ 360 QQQAPAGSAG LSAGAGAPGA SNGGFSLADS IARNIAINMA QWAQAQVQVP DAAPAPAGAP 420 DRQTVEAVAA AAAAAAAAAA TAVISAAGEA IQQRIQEKAA AGYLPFLVHP LAELIPPSLA 480 ARSGSAATTS DQSAPPWALP VPQQSLMHVL AGSVSTGGGA AGTHPHSAPP VSSRQQQQQQ 540 QQQQQQQQQQ QQQQQQQQQQ QTAPFSLGAA SGGAAGNAEQ QLTATRDDTH TGQHTGGTGG 600 TVTATPGSSA QQQGGAVSGP GNSPQHMSAS FAAVAAGGGS GGVASVSMAG GLGRLPSSSL 660 VASLGGPSVS LGGAAAMSLG VDLQPPQLLA PVASPPRMEW EVAVTVARAS QQQLQLQLQQ 720 QQQQQQQRQT APGFGLLAPQ PMAREGSAAA GLLRTLSLPA AWDSGGAAAQ AAAALQSREV 780 LSRLAAAYTS DVPQLQAVVA ATAAQQQGLP QGLLQPQPQL LQLVEQLPSR PSVELAQSPV 840 TLPGGGGSSS AFQVLRPATS QQQLEVSLDP RVRAVVDPRL QPQQQPRHVR SQSAAPGGAQ 900 PLPQARAEAQ GQQEAPAQAQ AQAQAQQEAQ AQAQAQAPYI KAEVLTEDEE RGRERERDPH 960 RHHHHHHHHA HHHHHHHSHH GSNHGGTYGG AHSRGIGSSA EPERGSRNPV DRRSRSVASI 1020 RSGGGCEERQ MHSGSGGGAG GNGLTRIGAA AAAAAAAVAA AAEEAAAQEQ VDKSSGASGC 1080 DNGSPAVSPD PRQLPLAVRG HGISPSAGAE GQLAHAGGGG TTGARVRFEA HVRSSTSEGR 1140 AHTGQQPQAT AAAAVAAAAV AGAGHRATAH RHGVHSSHRH RSSGGRGSAA AAAAATAAAS 1200 GAMDTDASAE PGGGARHRAG SRAAWLMDDY GAAASGDGAY QLGQLQYLPQ QFMPQQSGPD 1260 NGGLSGTTGS AFNSLQEVLQ GPDPAAMVPL LAPTSGVGAC GVQDAATTAQ LARLQLGAAH 1320 RSQAAERRVA AAERAAAAER AADGEGQEGG SGGNGVTDVN TYIRGDYSLL SPSGGNGGSG 1380 GYAAQLAGAV ATAAGSGGGG GSGGGSGGGG VPLSSSGPSS VPSTQQQGAN GGAATAMAAH 1440 RHSAQQTKQR AAAAAAAAAG AGGSDGNGAG ASNEMQQGSN QNAGSGAGSG QAGEAAGGVA 1500 QATGVALALP PPLPRSLLHV AARQGAPEAP SAERADAVGQ GACGGAAGLN AVSPTAGQPG 1560 GGSNTGSGSG DGSGAGGPPD VYQSSGPTGL ARAAAHAAGL RQETPPTGVS AGTRRGAGGS 1620 GSGNADGGGG ATGSGSGAGQ GSNPSGNGAG EGSGSGAAPA AGTAAGAANT GGVGSTQAAA 1680 GPHALQHPHA HGNAHQQQLQ QVQLQQQQQQ QQQGSGSGGE RGGSGSGAGS GSRVGSGVPA 1740 GTYAAREAMP GASGIGRHVP PYVVAAQRAQ HERVEAAAAK AAAATVGRDG EAGQADGSKP 1800 ISSPEADAEG ALLGSGGGSG GGAVVDLAVA LRHKQRGSPG GSGGMMGSQA RGDSPAAALT 1860 DPRLLQRGGS ALALAAGAAG RVASPPPNGQ STLPVPGDTL ALLHQMLSQQ QQQLQQAQAQ 1920 VVQQQQAQAA AAAAAGQDGG AAAVGSLAPM APPPHGLMTT LAGLSAGTQL AATLLALHEQ 1980 QQQQQQQEQL LATNRGGGSA AAGAGRSNGF LSLAAAAADP TSSGGGSSAT APPPLMALGP 2040 GALTALLPYL TGERPVESEH VARLEEVARA LQYASSLAGP LAAVLHPPGT SGPPAAQLAS 2100 ARDGLPSSDL AAAALMAQWQ NAATMYGLQP LLRTSSAAAA AAAAPGSATS GGSAAAPPPP 2160 SSAAAAAALL GFPPHPLLIQ QHHLQQQLAA SGQAQAAASY LQSLGLASGL GLGFNMAANM 2220 GSRSALGALS LGPMGLSLPT FASGGGGGNG TASGGALGAA AAAGGGGGAE QSPRPSAGTI 2280 GSGGGVHGLM PAPTRSHHQQ HSHQQHQQAA GAPSDPRLSG GNPRRGSGDG AVGFKPPRPL 2340 ASLGGVDGSG GVARGLGSGP SMSGEKLAAA AAAAAAAGGN GGTSKRSLNE LYGALVAAGA 2400 GAGGAAGGGG GGSGQGVQRS GRQRNGRRRH HRGHSPTRSV GSGASSSGFH CGDKSGGEEA 2460 NYGPRPSREY KEGSPQGGGA VEQQDTATSS GREEQGSAGA ARGCGAAGEV GTAGQSGGGA 2520 GGGAAAANAA AAANAAADGG GGMALPPAKH ARHDLMTGPN GKLAAAAGHG GGAGGASGQG 2580 TGGAAAKAAA QRPPKDATPS SGGGSGGSGG GSTGGNKSGT SNDAYDGSMN ASDEGSNEPG 2640 SAPADGPMGQ ALAPSDGMGM LGAGDPGLNV QGMQMPGAGM MGAGGARSFS VAQPEPGPGH 2700 ASIMAAQPGG CGGGGSTGVA GAGAGAGGGA AGGGGGTAGH KRVRYDAGPA GGGTAAAAGA 2760 PAAGAPLTPA AAGAAGAMQQ RAGALQADGA GTGTSSPSET GGGDDSRGAA QPPHKVQRTN 2820 AGGAAAVAEA GAGGAAGHAA DACDGNNRSG GSAGAGSGQA QSGRGRFAAP PSNQQHPHGA 2880 AAAAAAAGGA GGGAAQRTSG SLPSHLGYVL EQASSKQGGN DGSGNNNSGT GPRSGVGSNE 2940 GGAAGGAGGS RSGEGGNGNG SQGNGGSHGH GGSHGNGQGS NANGSHGHGS NGNNGHGSNG 3000 NGASTNLPGG APYAYFRTPL SLQQQPRGGG GGGGGGGGGG AGSGGSGGSG GNRGGGAQGS 3060 NGGNGDGAGS GGHGSGAHDG ASGGAAGPSP YDAGAPTSGA AADHMHRHLH HHHFTHAHHH 3120 YGGGGGGGPQ EGTSMRTPLP LQPQQPQPQP QQQQQQQQQQ QQQAGTGAAN AGQGMALAAV 3180 AAAAAAAPAV AGTSGGAAVV AAAAAAAAAV SQQQPVHVEG QGQDGNEPML QDG* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 1395 | 1405 | SGGGGGSGGGS |
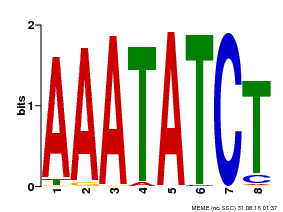
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00103 | PBM | Transfer from AT2G46830 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Cre12.g514400.t2.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A2K3D3N4 | 0.0 | A0A2K3D3N4_CHLRE; Uncharacterized protein | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G02840.3 | 9e-30 | LHY/CCA1-like 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cre12.g514400.t2.1 |



