 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | IGS.gm_5_00447 | ||||||||
| Common Name | CHLNCDRAFT_143118 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Trebouxiophyceae; Chlorellales; Chlorellaceae; Chlorella
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 427aa MW: 43740.4 Da PI: 8.8363 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 48.2 | 1.4e-15 | 34 | 64 | 1 | 31 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyr 31
C+ Cg+tkTp+WR+gp g ktLCnaCG++
IGS.gm_5_00447 34 CTKCGATKTPQWREGPFGAKTLCNACGVKRT 64
***************************9875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00401 | 4.8E-7 | 28 | 77 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 9.03E-11 | 31 | 66 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 1.5E-11 | 33 | 62 | IPR013088 | Zinc finger, NHR/GATA-type |
| Pfam | PF00320 | 2.8E-13 | 34 | 62 | IPR000679 | Zinc finger, GATA-type |
| CDD | cd00202 | 5.00E-11 | 34 | 79 | No hit | No description |
| PROSITE profile | PS50114 | 11.095 | 34 | 61 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 427 aa Download sequence Send to blast |
MTELAGPGAA HQSSPFDDEE FAFAHGAING VKCCTKCGAT KTPQWREGPF GAKTLCNACG 60 VKRTRKLRAE QEGAKRRRLS ASPAPPPQAK HAFAAPKYGA SRGAQYSQDA PTMDSYGSLE 120 PDSEAWGLPA AHGTRRPPRR AAEEAAFRTA RYARTGEWAE AQAHCEEAPR AATPSSSSDG 180 YSQPSSSCPE EVSWPPAAAG GAPRGSDCYA AINLMTMSAT QTAAAHAAAS CGAAPAAAAH 240 PCPAPAACPA SPASAALHFD AEQFNHDSLV ALAARYGAGA GAPPHHRVAP VTLSDLSRCL 300 PPAKVLELVR LNQELEVAVH EAHAASAAVA AVAQVLACKQ AAALRSRDVA GAATKRLRRF 360 MAELDTQFGI QSKFGAGRPP LSPTKPTGAA ALASPAKPLM AAARSPAAPV LAPLLPAAPA 420 LQARTL* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 74 | 78 | KRRRL |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00554 | DAP | Transfer from AT5G49300 | Download |
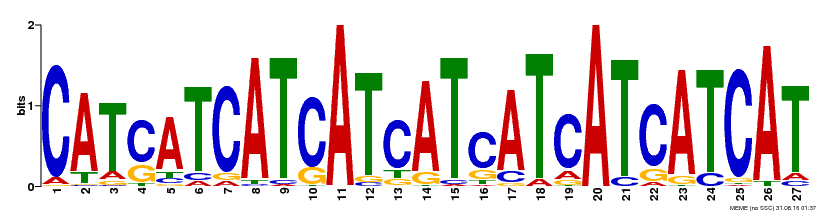
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_005849623.1 | 0.0 | hypothetical protein CHLNCDRAFT_143118 | ||||
| TrEMBL | E1Z9H7 | 0.0 | E1Z9H7_CHLVA; Uncharacterized protein | ||||
| STRING | XP_005849623.1 | 0.0 | (Chlorella variabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP220 | 14 | 30 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G36240.1 | 5e-12 | GATA transcription factor 7 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 17357301 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



