 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | IGS.gm_10_00281 | ||||||||
| Common Name | CHLNCDRAFT_133990 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; Trebouxiophyceae; Chlorellales; Chlorellaceae; Chlorella
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 294aa MW: 30469 Da PI: 4.6991 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 54.1 | 3.4e-17 | 147 | 208 | 1 | 62 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklkse 62
e+e kr rrkq+NRe+ArrsR+RK+ae+e+L + vk L +eN++Lk+e +l +++ l ++
IGS.gm_10_00281 147 EREMKRMRRKQSNRESARRSRLRKQAECEQLSRQVKDLASENSRLKEEKMQLLAQIEILNAK 208
799*********************************************99998888877655 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 5.2E-15 | 144 | 209 | No hit | No description |
| SMART | SM00338 | 1.9E-13 | 147 | 211 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 3.9E-15 | 147 | 207 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 12.013 | 149 | 212 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 8.84E-10 | 150 | 206 | No hit | No description |
| CDD | cd14702 | 7.97E-17 | 152 | 194 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 154 | 169 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 294 aa Download sequence Send to blast |
MGGFPLPGGE GEGKAEGQGS GEDETEEEEE EEDDGDSSGK EGGRPKRKRA ARASAKKKAA 60 TGDGVVKSNS NAALAMLAGA AATKPPNPAA AAAAAAALQQ NLGELWGTLN GAGAGGMPST 120 SGMDMSKLTS EQQQMLANAS FEGITDEREM KRMRRKQSNR ESARRSRLRK QAECEQLSRQ 180 VKDLASENSR LKEEKMQLLA QIEILNAKLS MSAFPGLHAM GGGDMTKQTA EIQAMAVAMA 240 QTMQGNGGVM MGTGLGIPTL DAKAMEGGDE VAAVPAAMEG AGEQQEQPEQ VSQ* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 163 | 169 | RRSRLRK |
| 2 | 163 | 170 | RRSRLRKQ |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00291 | DAP | Transfer from AT2G35530 | Download |
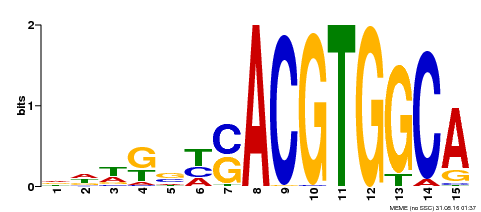
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_005847804.1 | 0.0 | hypothetical protein CHLNCDRAFT_133990 | ||||
| TrEMBL | E1ZER0 | 0.0 | E1ZER0_CHLVA; Uncharacterized protein | ||||
| STRING | XP_005847804.1 | 0.0 | (Chlorella variabilis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP4426 | 9 | 11 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G35530.1 | 9e-20 | basic region/leucine zipper transcription factor 16 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Entrez Gene | 17354908 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



