 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT2G35530.1 | ||||||||
| Common Name | AtbZIP16, bZIP16 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 409aa MW: 43109.3 Da PI: 5.2796 | ||||||||
| Description | basic region/leucine zipper transcription factor 16 | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 72 | 9e-23 | 303 | 365 | 1 | 63 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
++elkr+rrkq+NRe+ArrsR+RK+ae++eL++++++L +eN+ L+ e+++lk ++++l++e+
AT2G35530.1 303 DRELKRQRRKQSNRESARRSRLRKQAECDELAQRAEVLNEENTNLRAEINKLKSQCEELTTEN 365
689*********************************************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF07777 | 2.5E-28 | 1 | 103 | IPR012900 | G-box binding protein, multifunctional mosaic region |
| Pfam | PF16596 | 2.5E-46 | 142 | 272 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 1.2E-17 | 296 | 362 | No hit | No description |
| SMART | SM00338 | 1.8E-20 | 303 | 367 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 1.8E-20 | 304 | 365 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 13.173 | 305 | 368 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 1.5E-10 | 306 | 362 | No hit | No description |
| CDD | cd14702 | 1.88E-24 | 308 | 358 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 310 | 325 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 409 aa Download sequence Send to blast |
MASNEMEKSS KEKEPKTPPP SSTAPPSSQE PSSAVSAGMA TPDWSGFQAY SPMPPPHGYV 60 ASSPQPHPYM WGVQHMMPPY GTPPHPYVAM YPPGGMYAHP SMPPGSYPYS PYAMPSPNGM 120 TEVSGNTTGG TDGDAKQSEV KEKLPIKRSR GSLGSLNMIT GKNNEPGKNS GASANGAYSK 180 SGESASDGSS EGSDGNSQND SGSGLDGKDA EAASENGGSA NGPQNGSAGT PILPVSQTVP 240 IMPMTAAGVP GPPTNLNIGM DYWGAPTSAG IPGMHGKVST PVPGVVAPGS RDGGHSQPWL 300 QDDRELKRQR RKQSNRESAR RSRLRKQAEC DELAQRAEVL NEENTNLRAE INKLKSQCEE 360 LTTENTSLKD QLSLFPPLEG ISMDNDHQEP DTNQTGAAER KVDSYKDST |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 319 | 325 | RRSRLRK |
| 2 | 319 | 326 | RRSRLRKQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.37663 | 0.0 | seed| silique | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 145361658 | 0.0 | ||||
| Genevisible | 266634_at | 0.0 | ||||
| Expression Atlas | AT2G35530 | - | ||||
| AtGenExpress | AT2G35530 | - | ||||
| ATTED-II | AT2G35530 | - | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| TAIR | Encodes a G group bZIP transcription factor family member that can bind cis elements with an ACGT core, such as G-box, Hex, C-box and As-1. The protein is localized in the nucleus and can homodimerize and can heterodimerize with other G group members. | |||||
| UniProt | Transcriptional activator that binds to the G-box motif (5'-CACGTG-3') and other cis-acting elements with 5'-ACGT-3' core, such as Hex, C-box and as-1 motifs. Possesses high binding affinity to G-box, much lower affinity to Hex and C-box, and little affinity to as-1 element (PubMed:18315949). G-box and G-box-like motifs are cis-acting elements defined in promoters of certain plant genes which are regulated by such diverse stimuli as light-induction or hormone control (Probable). Binds to the G-box motif 5'-CACGTG-3' of LHCB2.4 (At3g27690) promoter. May act as transcriptional repressor in light-regulated expression of LHCB2.4. Binds DNA as monomer. DNA-binding activity is redox-dependent (PubMed:22718771). {ECO:0000269|PubMed:18315949, ECO:0000269|PubMed:22718771, ECO:0000305|PubMed:18315949}. | |||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
|
||||||
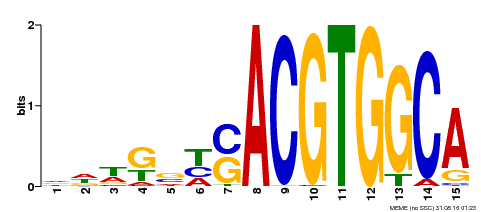
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00291 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT2G35530.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| BioGRID | AT4G36730, AT1G32150 | |||||
| IntAct | Search Q501B2 | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT2G35530 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT022055 | 0.0 | BT022055.1 Arabidopsis thaliana At2g35530 gene, complete cds. | |||
| GenBank | BT026441 | 0.0 | BT026441.1 Arabidopsis thaliana At2g35530 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_850248.2 | 0.0 | basic region/leucine zipper transcription factor 16 | ||||
| Swissprot | Q501B2 | 0.0 | BZP16_ARATH; bZIP transcription factor 16 | ||||
| TrEMBL | A0A178VPW6 | 0.0 | A0A178VPW6_ARATH; BZIP16 | ||||
| STRING | AT2G35530.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3701 | 25 | 57 | Representative plant | OGRP700 | 17 | 67 |
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT2G35530.1 |
| Entrez Gene | 818118 |
| iHOP | AT2G35530 |
| wikigenes | AT2G35530 |



