 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C023076P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 289aa MW: 31113.7 Da PI: 6.7458 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 56.2 | 4.8e-18 | 211 | 245 | 1 | 35 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkgl 35
C +Cg+ kTp+WR gp g+ktLCnaCG+++++ +l
MELO3C023076P1 211 CLHCGAEKTPQWRAGPFGPKTLCNACGVRFKSGRL 245
99*****************************9885 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF016992 | 2.7E-68 | 1 | 283 | IPR016679 | Transcription factor, GATA, plant |
| PROSITE profile | PS50114 | 11.793 | 205 | 241 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 3.1E-16 | 205 | 255 | IPR000679 | Zinc finger, GATA-type |
| SuperFamily | SSF57716 | 2.0E-14 | 207 | 269 | No hit | No description |
| Gene3D | G3DSA:3.30.50.10 | 3.9E-14 | 209 | 243 | IPR013088 | Zinc finger, NHR/GATA-type |
| CDD | cd00202 | 2.28E-13 | 210 | 258 | No hit | No description |
| Pfam | PF00320 | 5.1E-16 | 211 | 245 | IPR000679 | Zinc finger, GATA-type |
| PROSITE pattern | PS00344 | 0 | 211 | 236 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 289 aa Download sequence Send to blast |
MESSLAFMDD LLDFSSDIGE EDEEDDAVPP FSVKSKSSST TAPDSSDLNA AAMHPDDSSS 60 CRVLPEEDYA EEELEWLSNE DAFPAVETFV DILSDHHHHH APQPPPLTSV SKQNSPVSVL 120 ESTSISSHGE TINGGNKTSV HGSSILMSCC GGLKVPGKAR SKRRRGRHIS GHHLWFKQQP 180 SSKNLKQVVP TTETAAAVAA TTGAAGIGRK CLHCGAEKTP QWRAGPFGPK TLCNACGVRF 240 KSGRLVPEYR PASSPTFSAD LHSNSHRKVM EMRRQKQLGM VVNPMDKG* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that specifically binds 5'-GATA-3' or 5'-GAT-3' motifs within gene promoters. May be involved in the regulation of some light-responsive genes. {ECO:0000269|PubMed:12139008}. | |||||
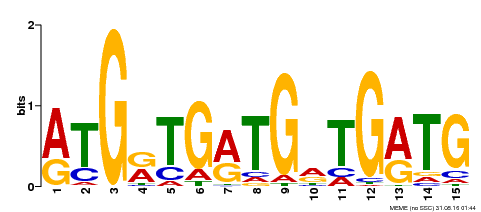
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00377 | DAP | Transfer from AT3G24050 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681844 | 0.0 | LN681844.1 Cucumis melo genomic scaffold, anchoredscaffold00057. | |||
| GenBank | LN713259 | 0.0 | LN713259.1 Cucumis melo genomic chromosome, chr_5. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008460721.1 | 0.0 | PREDICTED: GATA transcription factor 1 | ||||
| Swissprot | Q8LAU9 | 2e-61 | GATA1_ARATH; GATA transcription factor 1 | ||||
| TrEMBL | A0A1S3CCM6 | 0.0 | A0A1S3CCM6_CUCME; GATA transcription factor | ||||
| STRING | XP_008460721.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8537 | 29 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G24050.1 | 2e-53 | GATA transcription factor 1 | ||||



