 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MELO3C007540P1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Cucumis
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 369aa MW: 41348.5 Da PI: 7.3491 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.8 | 2.1e-32 | 243 | 298 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rFv+a+++LGGs++AtPk+i+elm+v+gLt+++vkSHLQkYRl+
MELO3C007540P1 243 KQRRCWSPELHRRFVDALHRLGGSQVATPKQIRELMQVDGLTNDEVKSHLQKYRLH 298
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.192 | 240 | 300 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.84E-18 | 240 | 301 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.7E-29 | 242 | 302 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.6E-26 | 243 | 298 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 2.3E-8 | 245 | 296 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010092 | Biological Process | specification of organ identity | ||||
| GO:0010629 | Biological Process | negative regulation of gene expression | ||||
| GO:0048449 | Biological Process | floral organ formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 369 aa Download sequence Send to blast |
MELSLDLSLD HSPKTIPQIL HELSSISDSF TKRSKLDDYV KRLEDEMRKI DAFKRELPLC 60 VLLLQDAILR LKDEVLQFKE QPVIEEFIPS KGISDQADEE NVKRKTRKWL SSAQLWSTNF 120 NFVDEEIANP KSNMNLDGDE DDRSVPQTPI ENWNCAKRRR AFELFKDQNN FVKRATNEDV 180 AFSEVPKLTL MTPISDPFPV NLTVKNGGNG GRNGRAAVSG LSSPAGQMKG QAKLSQQQQT 240 IRKQRRCWSP ELHRRFVDAL HRLGGSQVAT PKQIRELMQV DGLTNDEVKS HLQKYRLHVR 300 KLSPAEGSSG ENELKTSVTH AGSPDGPLHG GGSGKALSTT EGESMEVEED AKSDGHSWKG 360 RIQKHGDM* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 7e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 7e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 5e-14 | 243 | 296 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 5e-14 | 243 | 296 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 5e-14 | 243 | 296 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 5e-14 | 243 | 296 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 7e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 7e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 7e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 7e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 7e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 7e-14 | 243 | 296 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00629 | PBM | Transfer from PK14402.1 | Download |
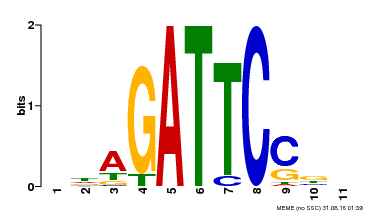
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681875 | 0.0 | LN681875.1 Cucumis melo genomic scaffold, anchoredscaffold00007. | |||
| GenBank | LN713262 | 0.0 | LN713262.1 Cucumis melo genomic chromosome, chr_8. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008440244.1 | 0.0 | PREDICTED: uncharacterized protein LOC103484758 | ||||
| TrEMBL | A0A1S3B0P5 | 0.0 | A0A1S3B0P5_CUCME; uncharacterized protein LOC103484758 | ||||
| STRING | XP_008440244.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1208 | 34 | 107 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G37180.1 | 1e-61 | G2-like family protein | ||||



