 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla022450 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | SRS | ||||||||
| Protein Properties | Length: 341aa MW: 36842.7 Da PI: 7.0514 | ||||||||
| Description | SRS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF702 | 226 | 7.3e-70 | 109 | 266 | 3 | 153 |
DUF702 3 sgtasCqdCGnqakkdCaheRCRtCCksrgfdCathvkstWvpaakrrerqqqlaaasskaaasa...aeaaskrkrel......kskkqsalsstklss 93
+g+++CqdCGnqakkdCa+ RCRtCCksrgf+C+thvkstWvpaakrrerqqq aa+++++++++ ++skr+re+ +++++sa+++ +l+s
Cla022450 109 AGGMNCQDCGNQAKKDCAYLRCRTCCKSRGFQCQTHVKSTWVPAAKRRERQQQIAALQQQQQQQEqfrGGENSKRQRETqtaaavTAATASASACPRLPS 208
6899****************************************************99998888766655566666665333322223333333344444 PP
DUF702 94 aeskkeletsslPeevsseavfrcvrvssvddgeeelaYqtavsigGhvfkGiLydqGle 153
+ + le +++P+ev+s+avfrcv+vs++dd++ee+aYqtav+igGhvfkGiLydqG+e
Cla022450 209 IT--SGLELAQFPSEVNSPAVFRCVKVSAMDDADEEFAYQTAVNIGGHVFKGILYDQGPE 266
43..3466677***********************************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05142 | 2.1E-68 | 111 | 266 | IPR007818 | Protein of unknown function DUF702 |
| TIGRFAMs | TIGR01623 | 4.4E-27 | 113 | 155 | IPR006510 | Zinc finger, lateral root primordium type 1 |
| TIGRFAMs | TIGR01624 | 8.7E-27 | 217 | 265 | IPR006511 | Lateral Root Primordium type 1, C-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009299 | Biological Process | mRNA transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 341 aa Download sequence Send to blast |
MAGFFYLGGR EGPPNKQEEE EEREQNLFLY RNEEIYNKGF EIWPQQYNNN NNHQQQQPNP 60 SLTNHLSFGV GPSRRSFMIN DPSDEPSRSA FTVMRPAAAA FAAASAATAG GMNCQDCGNQ 120 AKKDCAYLRC RTCCKSRGFQ CQTHVKSTWV PAAKRRERQQ QIAALQQQQQ QQEQFRGGEN 180 SKRQRETQTA AAVTAATASA SACPRLPSIT SGLELAQFPS EVNSPAVFRC VKVSAMDDAD 240 EEFAYQTAVN IGGHVFKGIL YDQGPEIRYT AAGESSSGHG GDGSTHPLNL IAGPTSATTS 300 AATSTNPSAA AAAAPLLDPS IYPAPLNAFM AGTQFFPPSR S |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds DNA on 5'-ACTCTAC-3' and promotes auxin homeostasis-regulating gene expression (e.g. YUC genes), as well as genes affecting stamen development, cell expansion and timing of flowering. Synergistically with other SHI-related proteins, regulates gynoecium, stamen and leaf development in a dose-dependent manner, controlling apical-basal patterning. Promotes style and stigma formation, and influences vascular development during gynoecium development. May also have a role in the formation and/or maintenance of the shoot apical meristem (SAM). Regulates anther dehiscence and floral development. {ECO:0000269|PubMed:16740146, ECO:0000269|PubMed:20706774}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00152 | DAP | Transfer from AT1G19790 | Download |
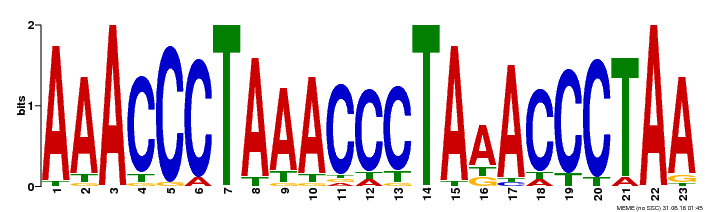
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681823 | 0.0 | LN681823.1 Cucumis melo genomic scaffold, anchoredscaffold00014. | |||
| GenBank | LN713257 | 0.0 | LN713257.1 Cucumis melo genomic chromosome, chr_3. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_023546679.1 | 1e-173 | protein SHI RELATED SEQUENCE 1-like | ||||
| Swissprot | Q9FXH7 | 9e-90 | SRS7_ARATH; Protein SHI RELATED SEQUENCE 7 | ||||
| TrEMBL | A0A0A0LNQ5 | 1e-154 | A0A0A0LNQ5_CUCSA; Uncharacterized protein | ||||
| STRING | XP_004173338.1 | 1e-155 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF882 | 34 | 125 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G75520.1 | 4e-74 | SHI-related sequence 5 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



