 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla015003 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 306aa MW: 33831.4 Da PI: 5.5641 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 86.2 | 2.9e-27 | 114 | 173 | 3 | 59 |
-SS-EEEEEEE--TT-SS-EEEEEE-ST...T---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 3 DgynWrKYGqKevkgsefprsYYrCtsa...gCpvkkkversaedpkvveitYegeHnhe 59
Dg++WrKYGqK++ gs+fpr+Y+rC+++ gC ++k+v++s++dp+v +itY+g+H+++
Cla015003 114 DGFSWRKYGQKDILGSKFPRGYFRCSHRftqGCLATKQVQKSDNDPTVYDITYKGRHTCN 173
9**************************9999***************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.20.25.80 | 1.4E-27 | 100 | 174 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 21.368 | 107 | 170 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 4.58E-24 | 109 | 174 | IPR003657 | WRKY domain |
| SMART | SM00774 | 1.0E-37 | 112 | 174 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 2.8E-25 | 114 | 172 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005730 | Cellular Component | nucleolus | ||||
| GO:0000987 | Molecular Function | core promoter proximal region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 306 aa Download sequence Send to blast |
MEFPTLIQEL NQGKQLANQL RNHLHPSSSS SSSSSSSSSS SSSSSHGILL IDQILRSYEN 60 ALLALSAGAP SVTSPLAPLN HTDVVSKKRK VMAKWSEQVK VSSSSANDSG PRCDGFSWRK 120 YGQKDILGSK FPRGYFRCSH RFTQGCLATK QVQKSDNDPT VYDITYKGRH TCNRVHSNTP 180 QEDQNSLLQH PMPPKQEQKP LQQPHDPSGF TFSSDAVQVK SENLDDVEGG LFQPFCAPSP 240 MFGSEVQDDQ SPFRESEFSP TFESNDMFGF CSDFETGFIS IPNSVTNISI GDLEEYCSFD 300 NVELFC |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5w3x_B | 3e-21 | 96 | 174 | 3 | 78 | Disease resistance protein RRS1 |
| 5w3x_D | 3e-21 | 96 | 174 | 3 | 78 | Disease resistance protein RRS1 |
| Search in ModeBase | ||||||
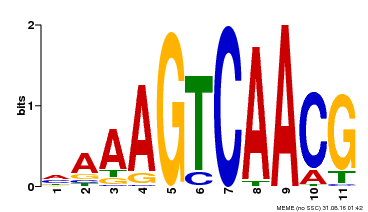
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00112 | ampDAP | Transfer from AT2G46400 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008461982.1 | 1e-170 | PREDICTED: probable WRKY transcription factor 53 | ||||
| TrEMBL | A0A1S3CGE2 | 1e-168 | A0A1S3CGE2_CUCME; probable WRKY transcription factor 53 | ||||
| STRING | XP_008461982.1 | 1e-169 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF24713 | 3 | 3 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46400.1 | 1e-40 | WRKY DNA-binding protein 46 | ||||



