 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla014058 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 317aa MW: 35714.9 Da PI: 7.1241 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 51.9 | 1.7e-16 | 14 | 62 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT.-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGgg.tWktIartmgkgRtlkqcksrwqkyl 48
+g+W++eEd +l+ +++q+G+g +W + ++++g++R++k+c++rw++yl
Cla014058 14 KGPWSPEEDAKLKAYIEQFGTGgNWIALPQKIGLKRCGKSCRLRWLNYL 62
79*********************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 37.7 | 4.7e-12 | 69 | 111 | 2 | 46 |
SSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 2 grWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
g +++eEd ++ + G++ W+ Ia+ ++ gRt++++k++w++
Cla014058 69 GGFSEEEDNIICSLYISIGSR-WSIIAAQLP-GRTDNDIKNYWNT 111
569******************.*********.***********97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.141 | 9 | 62 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.39E-26 | 11 | 109 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.1E-12 | 13 | 64 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.6E-15 | 14 | 62 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.0E-24 | 15 | 69 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.44E-10 | 16 | 62 | No hit | No description |
| PROSITE profile | PS51294 | 21.666 | 63 | 117 | IPR017930 | Myb domain |
| SMART | SM00717 | 4.8E-11 | 67 | 115 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-10 | 69 | 111 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-23 | 70 | 117 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 2.22E-8 | 71 | 113 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 317 aa Download sequence Send to blast |
MGRAPCCDKA NVKKGPWSPE EDAKLKAYIE QFGTGGNWIA LPQKIGLKRC GKSCRLRWLN 60 YLRPNIKHGG FSEEEDNIIC SLYISIGSRW SIIAAQLPGR TDNDIKNYWN TRLKKKLLGK 120 QQQALAARRI IHPIKKDDFD KNFNNNIIIH PAASVSSMNL HNYEAPYVQN PNTSTHDQDH 180 HHLKDLIMKM GGRFFCSDHH IPPSFQFNNP MDQFQSSNFG ELNMQFGVSN NDGTFHGVEQ 240 NSSGGEFGDY PIMLENMEDV NYGIMPSSST SGESSSLGEI SSLGFNYSDF EASQHHNIHH 300 VIAPAFADHH STNYSLQ |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 1e-24 | 14 | 117 | 7 | 108 | B-MYB |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00089 | SELEX | Transfer from AT3G49690 | Download |
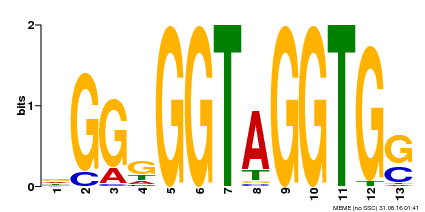
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KT885085 | 3e-86 | KT885085.1 Cucumis sativus cultivar TH169 1-aminocyclopropane-1-carboxylate synthase, hypothetical protein, DNA binding / transcription factor, and branched-chain amino acid aminotransferase genes, complete cds. | |||
| GenBank | KT885086 | 3e-86 | KT885086.1 Cucumis sativus cultivar WDSGD 1-aminocyclopropane-1-carboxylate synthase, hypothetical protein, DNA binding / transcription factor, and branched-chain amino acid aminotransferase genes, complete cds. | |||
| GenBank | KT885087 | 3e-86 | KT885087.1 Cucumis sativus cultivar WFEJ 1-aminocyclopropane-1-carboxylate synthase, hypothetical protein, DNA binding / transcription factor, and branched-chain amino acid aminotransferase genes, complete cds. | |||
| GenBank | KT885088 | 3e-86 | KT885088.1 Cucumis sativus cultivar WJE F11 1-aminocyclopropane-1-carboxylate synthase, hypothetical protein, DNA binding / transcription factor, and branched-chain amino acid aminotransferase genes, complete cds. | |||
| GenBank | KT885089 | 3e-86 | KT885089.1 Cucumis sativus cultivar TH118FLM 1-aminocyclopropane-1-carboxylate synthase, hypothetical protein, DNA binding / transcription factor, and branched-chain amino acid aminotransferase genes, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011657912.1 | 1e-173 | PREDICTED: transcription factor RAX2-like | ||||
| TrEMBL | A0A1S3B0P9 | 1e-167 | A0A1S3B0P9_CUCME; transcription factor RAX2 | ||||
| STRING | XP_008440439.1 | 1e-168 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF130 | 34 | 340 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G49690.1 | 2e-77 | myb domain protein 84 | ||||



