 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla012519 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 279aa MW: 30980.1 Da PI: 8.4932 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54.6 | 2.5e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT++Ed++l+d+++ +G g+W+t ++ g+ R++k+c++rw +yl
Cla012519 14 KGAWTKDEDQRLIDYIRVHGEGCWRTLPKAAGLLRCGKSCRLRWINYL 61
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 57.2 | 3.9e-18 | 67 | 111 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
rg++T++Edel+++++ +lG++ W+ Ia +++ gRt++++k++w+++
Cla012519 67 RGNFTEDEDELIIKLHSLLGNK-WSVIAGRLP-GRTDNEIKNYWNTH 111
89********************.*********.************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-23 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 16.454 | 9 | 61 | IPR017930 | Myb domain |
| SMART | SM00717 | 7.2E-13 | 13 | 63 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 4.21E-29 | 13 | 108 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 6.4E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.80E-10 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 27.908 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.8E-28 | 65 | 117 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 3.1E-17 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.1E-16 | 67 | 111 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.75E-13 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009611 | Biological Process | response to wounding | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 279 aa Download sequence Send to blast |
MGRSPCCEKA HTNKGAWTKD EDQRLIDYIR VHGEGCWRTL PKAAGLLRCG KSCRLRWINY 60 LRPDLKRGNF TEDEDELIIK LHSLLGNKWS VIAGRLPGRT DNEIKNYWNT HIKRKLITRG 120 IDPQTHRPLN ESAALSSSLP YRNHHHPTPN NSSSSSSPPP VAELPIMKPA INVHPSEPDE 180 EGSGTTTEMD PPLPAAEPMV AVHLKTEVNL ELSIGLHPFQ GEAARGSPGS SAESRLRQRS 240 PFICLCCQLG SEKGGECCRN CQRSYGWLYG FRPSALGSS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 3e-29 | 14 | 118 | 7 | 110 | B-MYB |
| Search in ModeBase | ||||||
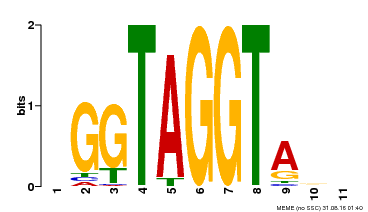
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00053 | PBM | Transfer from AT1G22640 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By ethylene, asbscisic acid (ABA), auxin (IAA), and Pseudomonas syringae pv. phaseolica. {ECO:0000269|PubMed:9839469}. | |||||
| UniProt | INDUCTION: By nitrogen, salicylic acid, NaCl and abscisic acid (ABA). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:18541146, ECO:0000269|PubMed:9839469}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004141649.1 | 1e-157 | PREDICTED: transcription repressor MYB6 | ||||
| Swissprot | Q38851 | 3e-92 | MYB6_ARATH; Transcription repressor MYB6 | ||||
| Swissprot | Q9S9K9 | 3e-92 | MYB3_ARATH; Transcription factor MYB3 | ||||
| TrEMBL | A0A0A0KXZ4 | 1e-156 | A0A0A0KXZ4_CUCSA; Uncharacterized protein | ||||
| STRING | XP_004141649.1 | 1e-157 | (Cucumis sativus) | ||||
| STRING | XP_004162864.1 | 1e-157 | (Cucumis sativus) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF264 | 34 | 221 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G22640.1 | 8e-93 | myb domain protein 3 | ||||



