 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla004431 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | WRKY | ||||||||
| Protein Properties | Length: 764aa MW: 82969.8 Da PI: 6.482 | ||||||||
| Description | WRKY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | WRKY | 101.7 | 4.4e-32 | 349 | 406 | 2 | 60 |
--SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS-- CS
WRKY 2 dDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhek 60
+DgynWrKYGqK+vkgse+prsYY+Ct+++C+vkkkvers+ +++++ei+Y+g Hnh+k
Cla004431 349 EDGYNWRKYGQKQVKGSEYPRSYYKCTHPNCQVKKKVERSH-EGHITEIIYKGAHNHPK 406
7****************************************.***************85 PP
| |||||||
| 2 | WRKY | 103.1 | 1.6e-32 | 564 | 622 | 1 | 59 |
---SS-EEEEEEE--TT-SS-EEEEEE-STT---EEEEEE-SSSTTEEEEEEES--SS- CS
WRKY 1 ldDgynWrKYGqKevkgsefprsYYrCtsagCpvkkkversaedpkvveitYegeHnhe 59
ldDgy+WrKYGqK+vkg+++prsYY+Ct +gC+v+k+ver+++d k v++tYeg+Hnh+
Cla004431 564 LDDGYRWRKYGQKVVKGNPNPRSYYKCTNPGCTVRKHVERASHDLKSVITTYEGKHNHD 622
59********************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.20.25.80 | 4.3E-28 | 347 | 407 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 9.29E-25 | 347 | 407 | IPR003657 | WRKY domain |
| SMART | SM00774 | 6.4E-36 | 348 | 406 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 23.158 | 349 | 407 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 1.2E-24 | 349 | 405 | IPR003657 | WRKY domain |
| Gene3D | G3DSA:2.20.25.80 | 6.6E-37 | 549 | 624 | IPR003657 | WRKY domain |
| SuperFamily | SSF118290 | 1.57E-28 | 556 | 624 | IPR003657 | WRKY domain |
| PROSITE profile | PS50811 | 38.124 | 559 | 624 | IPR003657 | WRKY domain |
| SMART | SM00774 | 7.5E-38 | 564 | 623 | IPR003657 | WRKY domain |
| Pfam | PF03106 | 8.5E-25 | 565 | 622 | IPR003657 | WRKY domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009555 | Biological Process | pollen development | ||||
| GO:0009942 | Biological Process | longitudinal axis specification | ||||
| GO:0030010 | Biological Process | establishment of cell polarity | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 764 aa Download sequence Send to blast |
MLRSSLDLGI RPHLGGFVVR ARDQVVMGRT DDNVAIIGDW VPPSPSPRTF FSAMQMLGED 60 IGSRPAMDTT SSDHKTEELF LRPREHTVSE NAVARGGIPG ANSGDRATEF GTFSEQKFRG 120 GLVERIAARA GFNAPRLNTE SIRSTDHSLN SEVKSPYLTI PPGLSPTTLL DSPVFLSNSL 180 AQQSPTTGKF QFLANVSNNR SSTMMSEANN NPFDDNSTSF AFRPSVESGS SFILSAAASK 240 TASATILPQS CPRIEVPVPR SENSFQSHLV EPSLSLPQNR IGHHPQVGLS TSYMEKDDVG 300 KTVSDDQRPF DSLCGGGEHS PPLDEQPDEG EQRGSGDSMA GSGCGAPSED GYNWRKYGQK 360 QVKGSEYPRS YYKCTHPNCQ VKKKVERSHE GHITEIIYKG AHNHPKPSPN RRVAIGSSDS 420 HINMQPDIPA QAAGQQSAEV SVWEDSQKGI PTGAPDWMQE NLEVTSSASL GPEYGNQPNP 480 LQAQNSSHIE TAEAIDASST FSNDEDEDDR GTHGSITLGY EGEGDESESK KRKLDAYVTE 540 MSGATRAIRE PRVVVQTTSE VDILDDGYRW RKYGQKVVKG NPNPRSYYKC TNPGCTVRKH 600 VERASHDLKS VITTYEGKHN HDVPAARNSS HISSGTSSPV TGQNSTAAMQ THVHRPGPPQ 660 PQNTIPRFER PAFGFAGRQQ MGTAFGMNQP GLGNLTMAAV GQAKLPVMPM HPYLAQAHHV 720 HEMGFLLPKG EPNVEPTSDL GLNFSNGSTV YQQIMSRLPL GPEM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wj2_A | 2e-37 | 349 | 625 | 8 | 78 | Probable WRKY transcription factor 4 |
| 2lex_A | 2e-37 | 349 | 625 | 8 | 78 | Probable WRKY transcription factor 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor. Regulates WOX8 and WOX9 expression and basal cell division patterns during early embryogenesis. Interacts specifically with the W box (5'-(T)TGAC[CT]-3'), a frequently occurring elicitor-responsive cis-acting element. Required to repolarize the zygote from a transient symmetric state. {ECO:0000269|PubMed:21316593}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00071 | PBM | Transfer from AT5G56270 | Download |
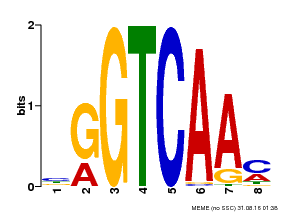
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | GU984016 | 0.0 | GU984016.1 Cucumis sativus WRKY protein (WRKY16) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008448990.1 | 0.0 | PREDICTED: probable WRKY transcription factor 2 | ||||
| Swissprot | Q9FG77 | 0.0 | WRKY2_ARATH; Probable WRKY transcription factor 2 | ||||
| TrEMBL | A0A1S3BLY1 | 0.0 | A0A1S3BLY1_CUCME; probable WRKY transcription factor 2 | ||||
| STRING | XP_008448990.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF9524 | 34 | 43 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G56270.1 | 0.0 | WRKY DNA-binding protein 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



