 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cla001961 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Cucurbitales; Cucurbitaceae; Benincaseae; Citrullus
|
||||||||
| Family | Trihelix | ||||||||
| Protein Properties | Length: 545aa MW: 61805.7 Da PI: 5.7195 | ||||||||
| Description | Trihelix family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | trihelix | 87.1 | 2.1e-27 | 58 | 143 | 1 | 86 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqle 86
rW++qe+laL+++r++m+ +r++ k+plW+evs+k+ e gf+r+pk+Ckek+en+ k++k++k+ +++++++s++ +++ d+le
Cla001961 58 RWPRQETLALLKIRSDMDAIFRDATHKAPLWDEVSRKLGELGFNRTPKKCKEKFENVYKYHKRTKDVRSGKSDNSKKVYRFSDELE 143
8*********************************************************************9999999999999987 PP
| |||||||
| 2 | trihelix | 105.7 | 3.2e-33 | 379 | 464 | 1 | 87 |
trihelix 1 rWtkqevlaLiearremeerlrrgklkkplWeevskkmrergferspkqCkekwenlnkrykkikegekkrtsessstcpyfdqlea 87
rW+k ev aLi++r+eme +++++ k+ lWee+s++mr g++rs+k+Ckekwen+nk++kk+k+++kkr +e+s+tcpyf+ql+a
Cla001961 379 RWPKGEVEALIRLRTEMEMKYQENGPKGLLWEEISAAMRGLGYNRSSKRCKEKWENINKYFKKVKDSNKKR-AEDSKTCPYFHQLDA 464
8*********************************************************************8.89999********85 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50090 | 7.364 | 51 | 115 | IPR017877 | Myb-like domain |
| SMART | SM00717 | 2.0E-4 | 55 | 117 | IPR001005 | SANT/Myb domain |
| CDD | cd12203 | 1.12E-22 | 57 | 122 | No hit | No description |
| Pfam | PF13837 | 3.8E-17 | 57 | 138 | No hit | No description |
| SMART | SM00717 | 1.7E-4 | 376 | 438 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-4 | 378 | 435 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50090 | 7.491 | 378 | 436 | IPR017877 | Myb-like domain |
| CDD | cd12203 | 5.15E-26 | 378 | 443 | No hit | No description |
| Pfam | PF13837 | 4.1E-23 | 379 | 465 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 545 aa Download sequence Send to blast |
MDVDSGEAVV TEAGEAHEVG NGDGRGGGGG NGSNSGEEDK GSSLLLFEDG EKNFGGNRWP 60 RQETLALLKI RSDMDAIFRD ATHKAPLWDE VSRKLGELGF NRTPKKCKEK FENVYKYHKR 120 TKDVRSGKSD NSKKVYRFSD ELEAFDHHNS HHQNHILFQS HHHQPPQPPP PLPPVPPTTT 180 TTPPPVMKTV NSTVSSTMNT TTNNNSLPPK SSNNPLSNLP NMAANVMFSS STSSSTASEE 240 DPFRSSRRRR KKRKWSDFFL RLTKEVIEKQ EGLQLKFLEA LERIENQRKL RDEAWRMKEM 300 KRVNQEHEVL VQEMSMAAAK DAAVVAFLQK IAPFSAPPPP PPPPPMTTQS QNNENNGKMT 360 SVIPMTNGKV SSIGGSPSRW PKGEVEALIR LRTEMEMKYQ ENGPKGLLWE EISAAMRGLG 420 YNRSSKRCKE KWENINKYFK KVKDSNKKRA EDSKTCPYFH QLDALYREKE KNMNNFDINS 480 QMEPLMVEPE QQWPPPFQAN NQIMGSLQRI NDEGNQEEDD DEEEEEEEEE EEEEDGGSSS 540 TDVED |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 247 | 252 | RRRKKR |
| 2 | 247 | 253 | RRRKKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that binds specific DNA sequence. {ECO:0000250}. | |||||
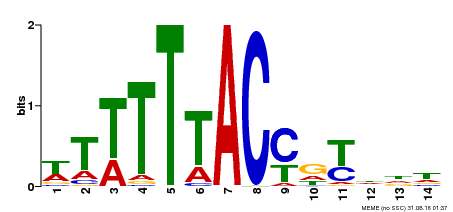
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00244 | DAP | Transfer from AT1G76890 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | LN681817 | 1e-177 | LN681817.1 Cucumis melo genomic scaffold, anchoredscaffold00015. | |||
| GenBank | LN713257 | 1e-177 | LN713257.1 Cucumis melo genomic chromosome, chr_3. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011656564.1 | 0.0 | PREDICTED: trihelix transcription factor GT-2-like | ||||
| Swissprot | Q39117 | 1e-124 | TGT2_ARATH; Trihelix transcription factor GT-2 | ||||
| TrEMBL | A0A0A0KE50 | 0.0 | A0A0A0KE50_CUCSA; Uncharacterized protein | ||||
| STRING | XP_008445762.1 | 0.0 | (Cucumis melo) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF373 | 34 | 181 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G76880.1 | 2e-70 | Trihelix family protein | ||||



