 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Cagra.1655s0043.1.p | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Capsella
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 265aa MW: 29626.1 Da PI: 7.2959 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 24.7 | 5.4e-08 | 33 | 79 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwqky 47
+WT+eE + + +a + + + +W ++a++++ g+t ++ + + k+
Cagra.1655s0043.1.p 33 SWTKEENKMFERALAIYAEDspdRWFKVASMIP-GKTVLDVMNQYSKL 79
7*****************99*************.*********99876 PP
| |||||||
| 2 | Myb_DNA-binding | 39.9 | 9.9e-13 | 129 | 173 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE+ +++ + ++G+g+W+ I+r + +t+ q+ s+ qky
Cagra.1655s0043.1.p 129 PWTEEEHRRFLLGLLKYGKGDWRNISRNFVVSKTPTQVASHAQKY 173
8*****************************99************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-4 | 28 | 78 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51293 | 9.449 | 29 | 82 | IPR017884 | SANT domain |
| SMART | SM00717 | 5.8E-9 | 30 | 82 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 1.15E-11 | 32 | 86 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 8.4E-7 | 33 | 78 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.67E-8 | 34 | 79 | No hit | No description |
| PROSITE profile | PS51294 | 19.439 | 122 | 178 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.51E-15 | 124 | 177 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.5E-17 | 125 | 177 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 6.4E-11 | 126 | 178 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.7E-10 | 126 | 176 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.3E-10 | 129 | 173 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.38E-9 | 129 | 173 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 265 aa Download sequence Send to blast |
METLHPLSHL PISDHRFVFQ EMMSLQSSSS SSSWTKEENK MFERALAIYA EDSPDRWFKV 60 ASMIPGKTVL DVMNQYSKLE EDVSDIEAGR VPIPGYPAAS SPSAFDPDTC RKRPNGARGS 120 DHDRKKGVPW TEEEHRRFLL GLLKYGKGDW RNISRNFVVS KTPTQVASHA QKYYQRQLSG 180 AKDKRRPSIH DITTVNLLKA NLNRSFSDHS DILPDLGFTD KDNAEEGVMF MGQNLSSETL 240 FSPSPTSFDA AFNFAGANAF SARA* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 4e-16 | 27 | 96 | 4 | 73 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00347 | DAP | Transfer from AT3G11280 | Download |
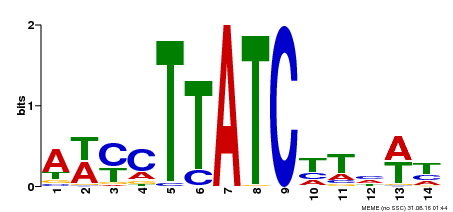
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Cagra.1655s0043.1.p |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY056180 | 0.0 | AY056180.1 Arabidopsis thaliana putative MYB-family transcription factor (At3g11280) mRNA, complete cds. | |||
| GenBank | AY091265 | 0.0 | AY091265.1 Arabidopsis thaliana putative MYB-family transcription factor (At3g11280) mRNA, complete cds. | |||
| GenBank | AY550308 | 0.0 | AY550308.1 Arabidopsis thaliana MYB transcription factor (At3g11280) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006298342.1 | 0.0 | transcription factor DIVARICATA isoform X2 | ||||
| Refseq | XP_023641495.1 | 0.0 | transcription factor DIVARICATA isoform X1 | ||||
| Swissprot | Q8S9H7 | 5e-74 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | R0I032 | 0.0 | R0I032_9BRAS; Uncharacterized protein | ||||
| STRING | Cagra.1655s0043.1.p | 0.0 | (Capsella grandiflora) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM4340 | 26 | 57 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G11280.2 | 1e-162 | MYB family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Cagra.1655s0043.1.p |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



